Sabitlenmiş Tweet

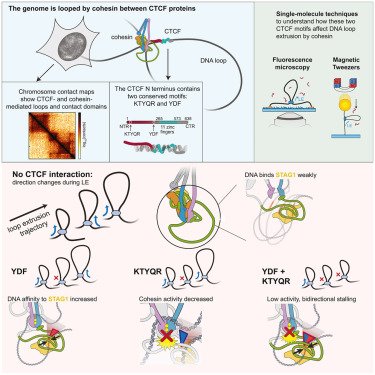
Exciting news 📃🧬🔬 Today we report on the dynamics of a fascinating class of molecular motors, so-called SMC proteins in @CellCellPress
doi.org/10.1016/j.cell…

English
Roman Barth
326 posts

@RomanBarth2
Postdoc at David Baker lab @ IPD U Washington | PhD at Cees Dekker lab @ TU Delft | Schmidt Science Fellow | Creating and understanding the tiniest machines