Sabitlenmiş Tweet

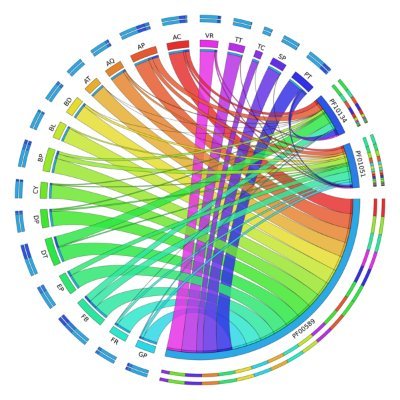
I’m very proud to announce my first publication. Featured on the cover of G3 Genes | Genomes | Genetics. The haplotype-resolved genome of European hazelnut will be a valuable resource for the plant breeding program here at Oregon State University.
academic.oup.com/g3journal/arti…

English