Kurt Schmoller
972 posts





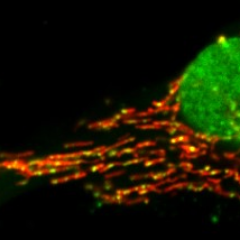
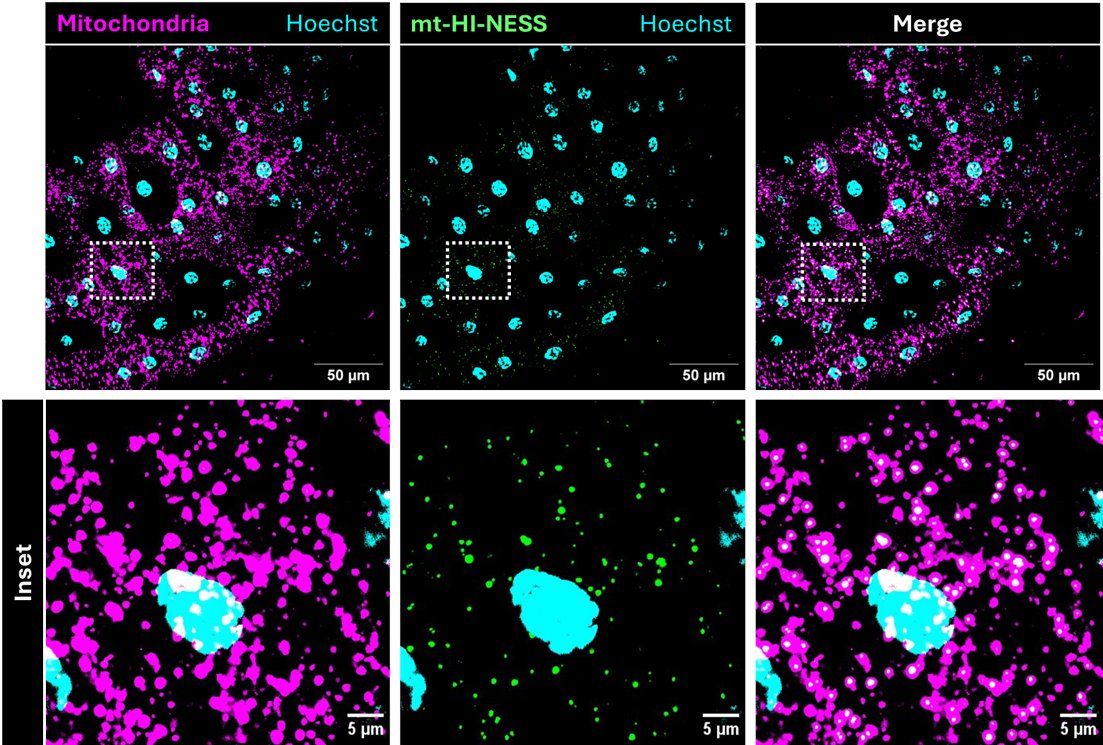
I want to follow up on our recent preprint introducing a new genetic fluorescent mtDNA nucleoid reporter. biorxiv.org/content/10.110…





Cover & highlight #22 by @osman_lab N&V on senescence/p21 & the path to fibrosis Commentary on PIWI & RNAs and… CEP192 localises mitotic Aurora-A activity DELE1 maintains muscle proteostasis Carboxy-terminal polyglutamylation regulates Dishevelled LLPS … embopress.org/toc/14602075/4…

Cover & highlight #22 by @osman_lab N&V on senescence/p21 & the path to fibrosis Commentary on PIWI & RNAs and… CEP192 localises mitotic Aurora-A activity DELE1 maintains muscle proteostasis Carboxy-terminal polyglutamylation regulates Dishevelled LLPS … embopress.org/toc/14602075/4…





🚀🎉SpotMAX is now a proud community partner of @bioimageio, the model zoo where you can find awesome pre-trained segmentation models! PS: You won't get the fireworks on the BioImage.IO website but feel free to request them on the SpotMAX GUI 😎

SpotMAX, our software tool for analysing multi-dimensional microscopy data, is finally out! 🎉 And conveniently, I just presented it at the #I2K conference 😀 A multi-year-long effort of many great collaborators and users. But what can SpotMAX do? A small thread






📢 JOB ALERT: Tvardovskiy Lab at @PennEpiInst is hiring at all levels! Interested in chromatin biology, quantitative proteomics, or epigenetic modifications? Learn more about our research and apply at tvardovskiylab.org Please RT




You want to analyze your cells and need to quantify spots? But you don't want to develop your own pipeline from scratch or click on spots manually? @frank_pado and #spotmax have you covered! doi.org/10.1101/2024.1…


SpotMAX, our software tool for analysing multi-dimensional microscopy data, is finally out! 🎉 And conveniently, I just presented it at the #I2K conference 😀 A multi-year-long effort of many great collaborators and users. But what can SpotMAX do? A small thread