Sabitlenmiş Tweet

Pleased to share my latest review on Cell-Free Protein Synthesis #CFPS and #AI/ML for accelerating drug discovery
sciencedirect.com/science/articl…
#CellFree #SynBio
English
Filippo Caschera
1.5K posts


@Stjle
Synthetic Biology, Artificial Cells, Amphiphile Systems, Machine Learning, Cell-Free Protein Synthesis: Italy, Denmark, Japan, USA, France, Germany, UK.






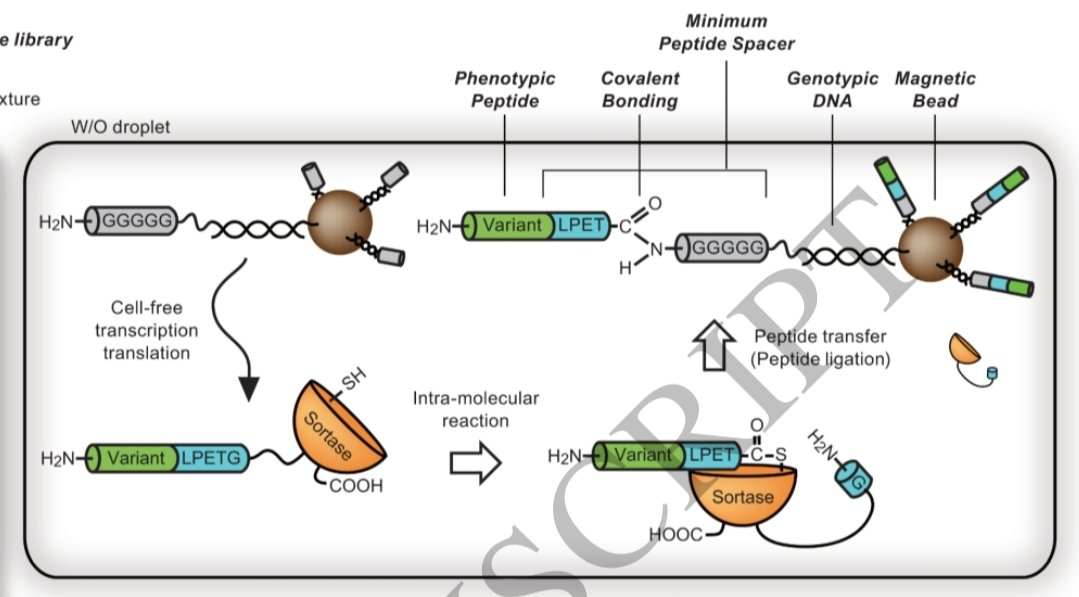
計画班・松浦グループの研究成果です。膜タンパク質を進化させる無細胞分子システムを構築し、新たな膜タンパク質変異体を造り出すことに成功しました。得られた変異体膜タンパク質が、対象膜タンパク質の生体組織での役割を解明するのに有用であることも示しました。pubs.acs.org/doi/10.1021/ja…