Streamlining large-scale genomic data management: Insights from the UK Biobank whole-genome sequencing data dlvr.it/TN8k2L
Tony
32 posts

@Tony_Chen6
Postdoc @MGHMedicine / @BroadInstitute || PhD from @HarvardBiostats




Streamlining large-scale genomic data management: Insights from the UK Biobank whole-genome sequencing data dlvr.it/TN8k2L






I'm excited to present at the Program in Quantitative Genomics @HarvardPqg Working Group seminar next Tuesday. It's always a pleasure to receive seminar invitations from students. Thank you, @Tony_Chen6. I'm looking forward to it!


Attending #ASHG24? Excited with new statistical genetics methods and analyzing data from newest functional genomics tech? Happy to chat if you are interested in joining my group @uwgenome UW in beautiful Seattle next spring or later 🙂


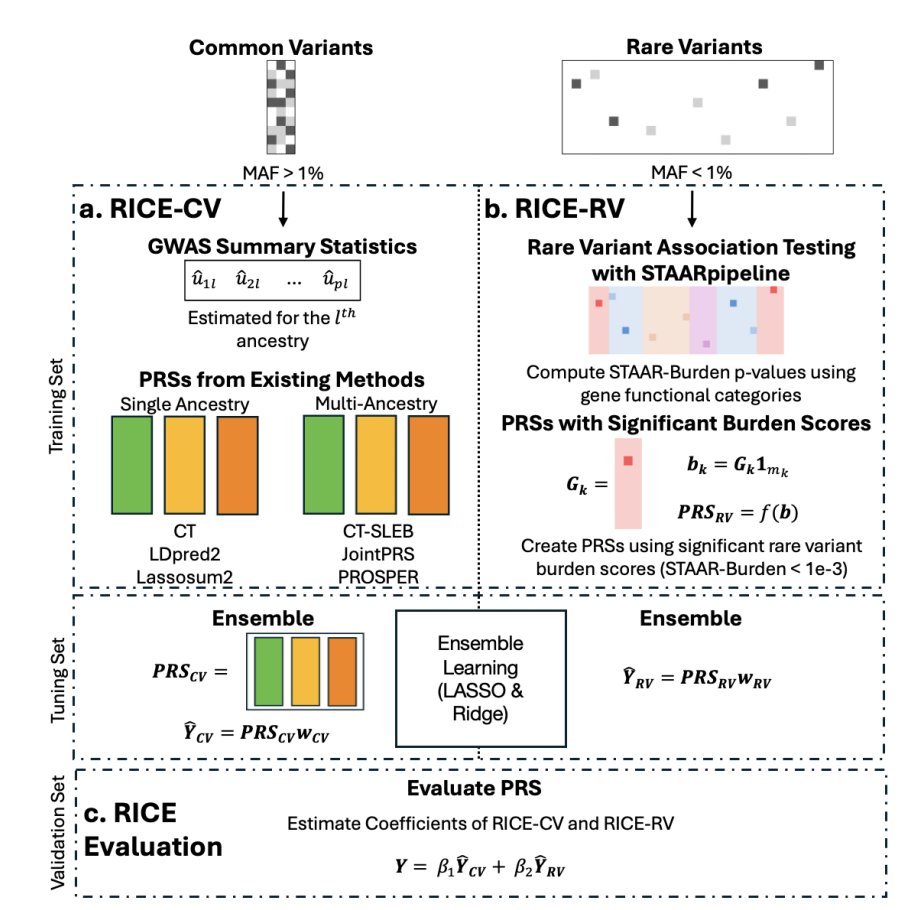
Very excited to share our preprint presenting a framework for translation of PRS to primary care for lung cancer in diverse high-risk patients. Grateful for the support from @AndrewHaoyu and @lishiunchen, and amazing collaborators from WashU and ILCCO. medrxiv.org/content/10.110…


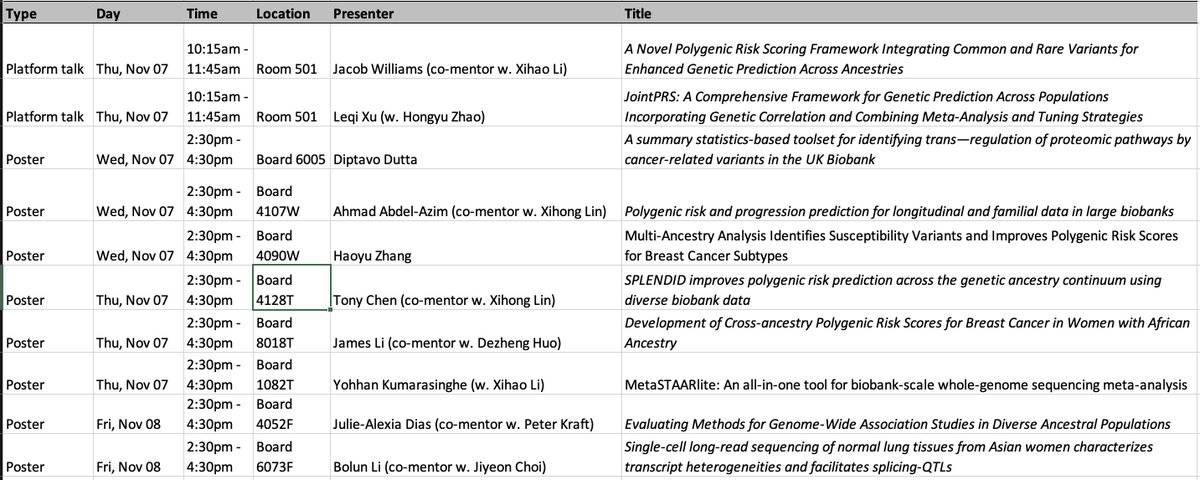



SPLENDID enables computation of a single, label-free model with robust prediction across all ancestries, which we hope can enhance fairness and interpretability in clinical implementation. Software and tutorials are available on GitHub. github.com/chen-tony/SPLE…

Using training data from All of Us and validation in UK Biobank, SPLENDID outperforms existing methods, particularly in non-European ancestry, selects fewer variants, remains robust to ancestry proportions in tuning, and can be used with only training data.

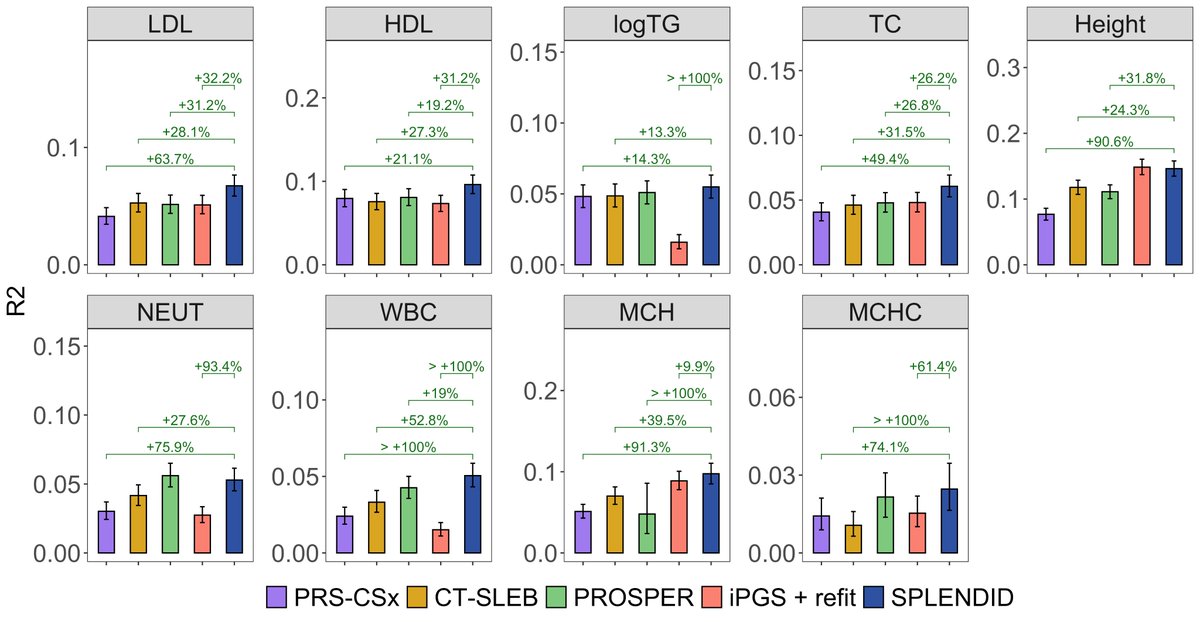
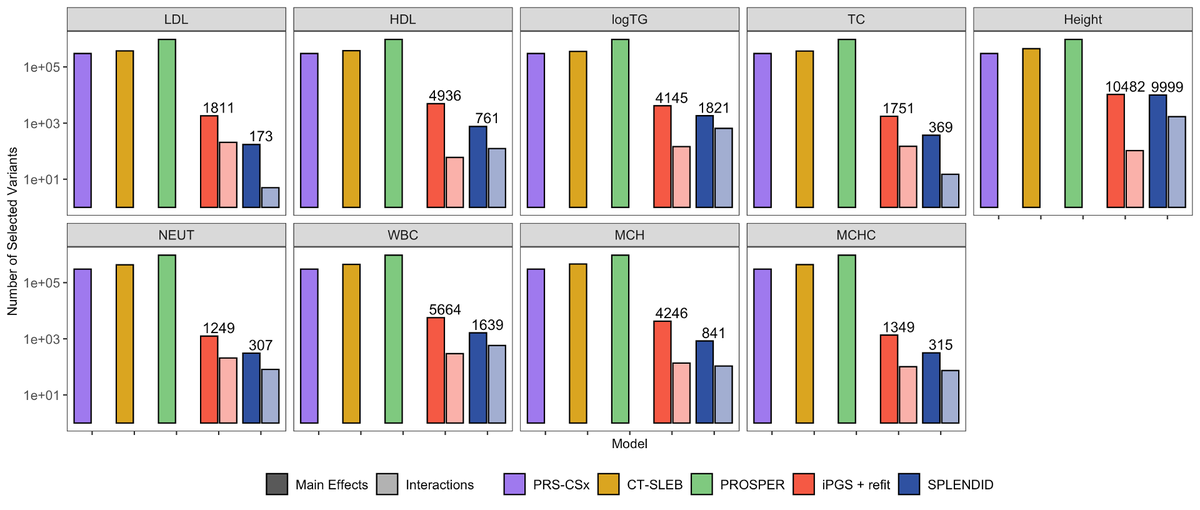
SPLENDID uses an ensemble group-L0L2 penalized regression framework to model interactions between genetic variants and ancestry PCs. This allows for simultaneous estimation of shared and heterogeneous effects across diverse populations without discrete ancestry labels.

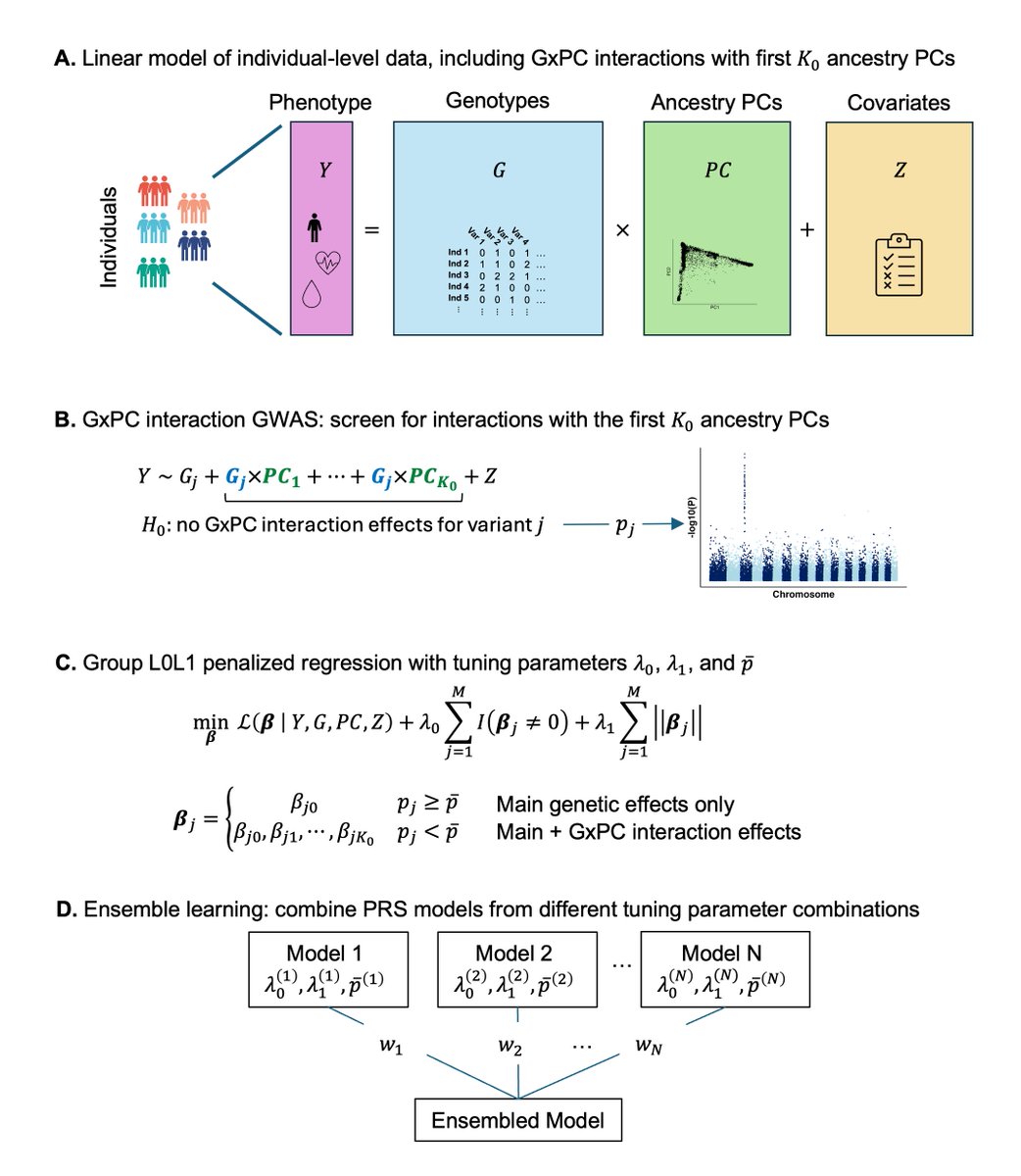
Excited to share our preprint for SPLENDID, a new PRS method using continuous ancestry interactions and biobank-scale individual level data. Many thanks to @AndrewHaoyu , Rahul Mazumder, and @XihongLin for their valuable guidance on this work! biorxiv.org/content/10.110…



Hi - I will give a seminar about our recently published Nature paper on “mosaic X chromosome loss” at the PQG working group NEXT TUESDAY! Welcome to Join the talk and discussion in person!! ⏰Time: 1-2 PM, Tuesday, Oct 22 🏠Location: Biostats Conference Room (2-426), Harvard Chan School (Thanks, @Tony_Chen6, for the invitation! Excellent seminars have also been scheduled for Nov 19 by @saorisakaue and Dec 17 by @yk_tani!)

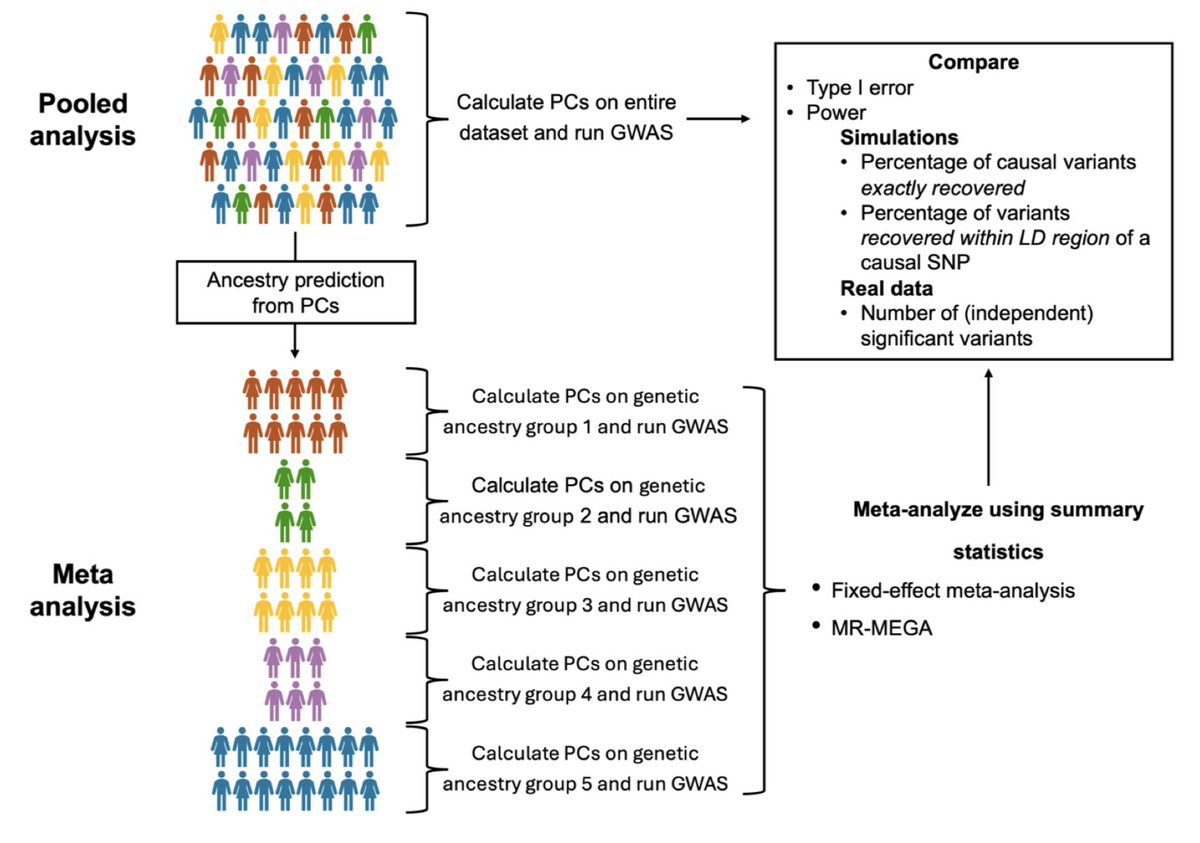
📢Online now! 📰Semi-supervised machine learning method for predicting homogeneous ancestry groups to assess Hardy-Weinberg equilibrium in diverse whole #genome sequencing studies 🧑🤝🧑@xihonglin @bmneale @mczody & co cell.com/ajhg/abstract/…



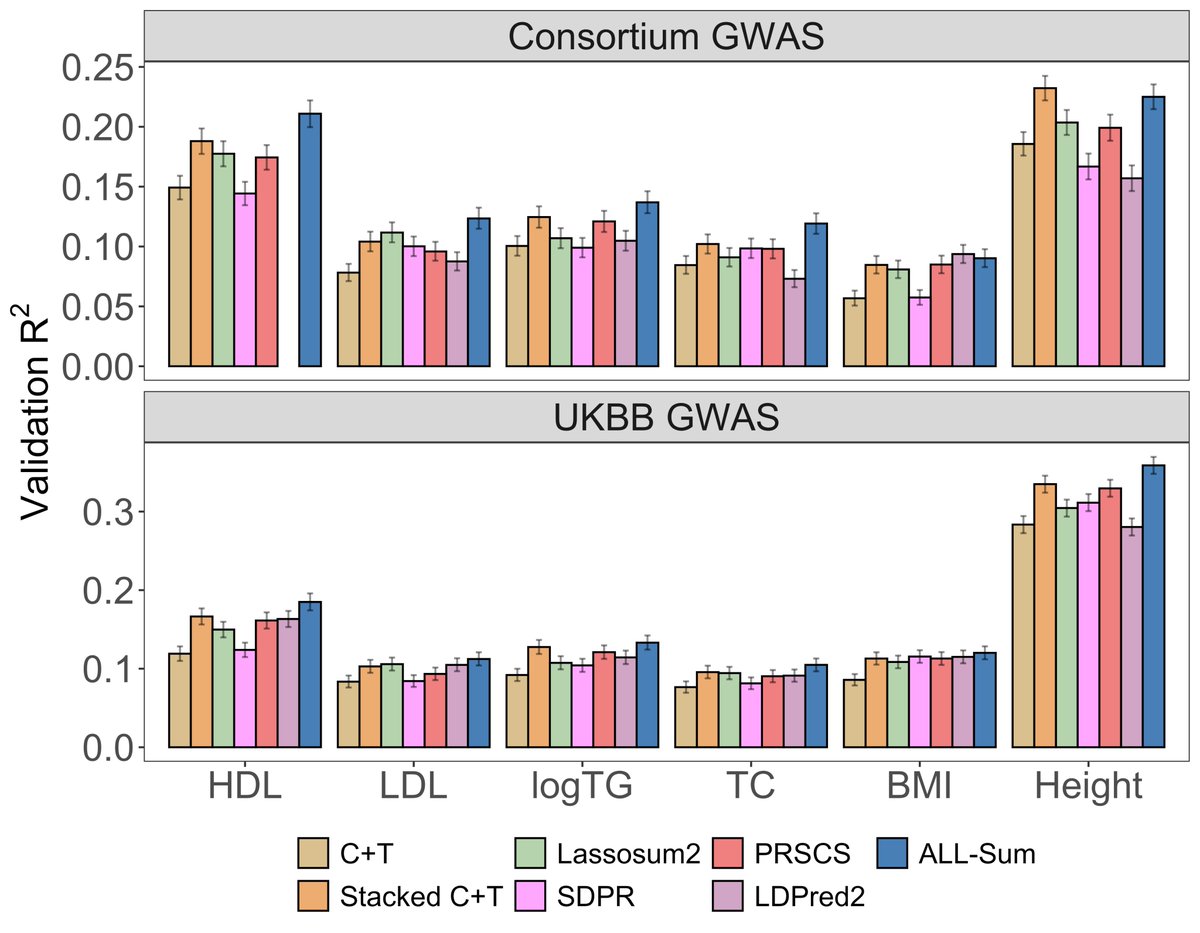
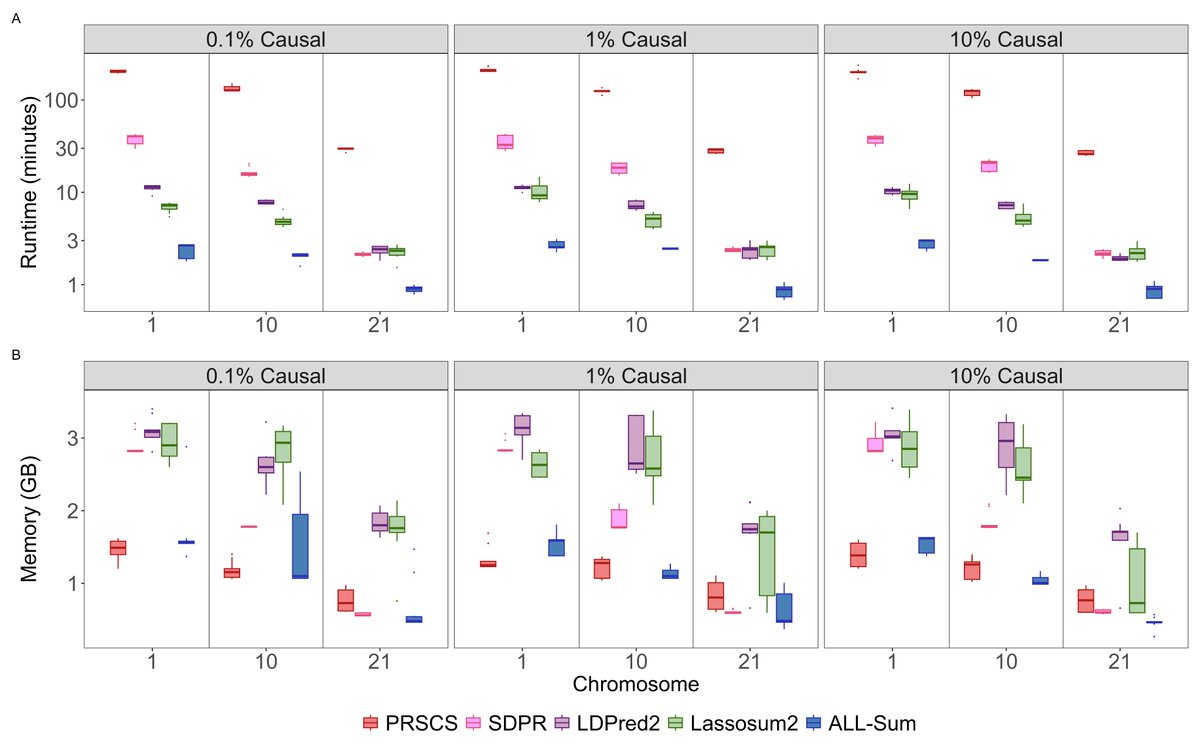
Our paper introducing a fast and scalable PRS method (ALL-Sum) is now available on @PNASNews! Many thanks to @AndrewHaoyu, Rahul Mazumder, and @XihongLin for their mentorship, as well as @BhramarBioStat and @HongyuZhao2 for reviewing! pnas.org/doi/10.1073/pn…