
Aaron Jesse O'Hare-Lewis 🏳️🌈
65 posts


@aaron_jesse_
Consultant @BCG, previously PhD @Cambridge_Uni @MRC_LMB, BS @Yale MB&B.
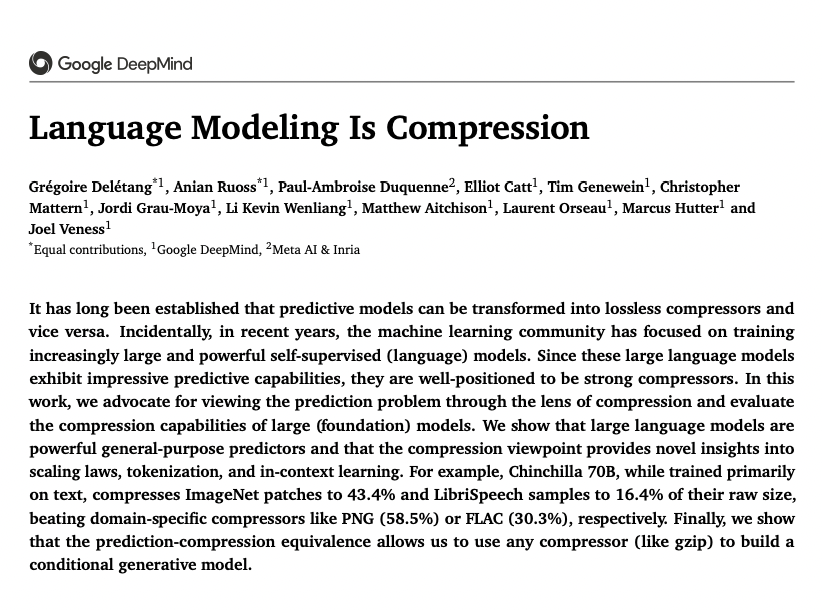














Language Modeling Is Compression paper page: huggingface.co/papers/2309.10… It has long been established that predictive models can be transformed into lossless compressors and vice versa. Incidentally, in recent years, the machine learning community has focused on training increasingly large and powerful self-supervised (language) models. Since these large language models exhibit impressive predictive capabilities, they are well-positioned to be strong compressors. In this work, we advocate for viewing the prediction problem through the lens of compression and evaluate the compression capabilities of large (foundation) models. We show that large language models are powerful general-purpose predictors and that the compression viewpoint provides novel insights into scaling laws, tokenization, and in-context learning. For example, Chinchilla 70B, while trained primarily on text, compresses ImageNet patches to 43.4% and LibriSpeech samples to 16.4% of their raw size, beating domain-specific compressors like PNG (58.5%) or FLAC (30.3%), respectively. Finally, we show that the prediction-compression equivalence allows us to use any compressor (like gzip) to build a conditional generative model.







Happy Birthday, Johann Gregor Mendel! Your groundbreaking work in genetics has paved the way for countless discoveries in the field. Your legacy continues to inspire generations of scientists today. 🫛🧬 #FatherOfGenetics #CzechRepublic #Science 20 July 1822 – 6 January 1884


