

Bin Zhang
162 posts

@binzmit
Associate professor of chemistry, MIT






I am excited to share our latest #preprint: ChromoGen: Diffusion model predicts single-cell chromatin conformations researchsquare.com/article/rs-463… As the title suggests, we achieved de novo prediction of 3D chromatin structures using DNA sequence and ATAC-seq using AI.

Protein Language Model Identifies Disordered, Conserved Motifs Driving Phase Separation 1. This study employs ESM2, a cutting-edge protein language model, to analyze intrinsically disordered regions (IDRs) in proteins, uncovering conserved motifs critical for phase separation and membraneless organelle (MLO) formation. 2. A major finding reveals that IDRs involved in phase separation contain conserved “sticker” residues (e.g., Y, W, F) and “spacer” residues (e.g., G, A, P), forming functional sequence motifs under evolutionary pressure. 3. By predicting mutational constraints, ESM2 accurately identifies conserved, functional residues and motifs without relying on sequence alignments, overcoming a key limitation in studying disordered proteins. 4. Experimental validation shows that many conserved motifs identified by ESM2 are essential for phase separation. Mutations within these motifs disrupt MLO formation, highlighting their biological significance. 5. The study introduces a motif-based framework for understanding the “grammar” of IDRs, emphasizing how conserved motifs integrate structural flexibility with functional specificity. 6. This work bridges computational predictions with experimental biology, advancing our understanding of IDRs in phase separation and offering insights into disease-linked protein dysfunction. 7. ESM2 emerges as a powerful tool for investigating the evolutionary and functional landscapes of disordered proteins, with broad implications for molecular biology and synthetic biology. @binzmit @yumengzhang99 📜Paper: biorxiv.org/content/10.110… #ProteinScience #Bioinformatics #PhaseSeparation #MachineLearning #IntrinsicallyDisorderedRegions

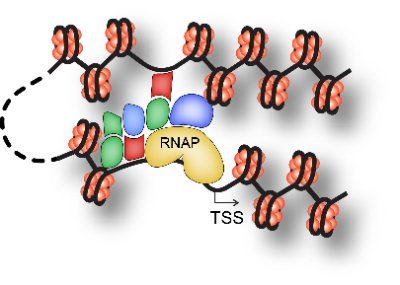







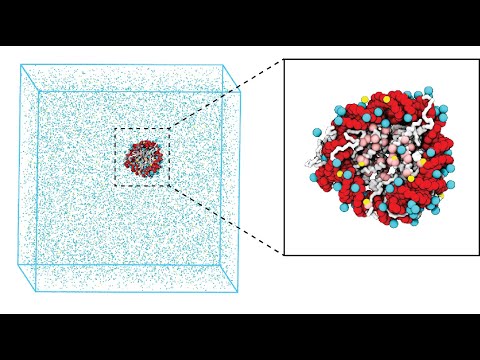
It's as if the very fabric of chromatin's existence is intricately woven through these inherent interactions, shaping its three-dimensional structure. biorxiv.org/content/10.110…


A new paper from Songtao Ye @ye_songtao44002 and Xin Zhang @kfzhangxin @Westlake_Uni reveals the critical role of micropolarity in controlling the structure and miscibility of subcompartments in #biomolecular condensates. Read it here rdcu.be/driFa


