
Our X-cell is up at @biorxiv_bioinfo ! Read our full paper at biorxiv.org/content/10.648… Part of the data and the model weights will be shared soon. stay tuned!
ChloeXWang
29 posts

@ChloeXWang1
CS PhD student at the WangLab @ U of T. Working on ML applications in single-cell research.

Our X-cell is up at @biorxiv_bioinfo ! Read our full paper at biorxiv.org/content/10.648… Part of the data and the model weights will be shared soon. stay tuned!


We @arcinstitute, @UHN, and @VectorInst recently released out BioReason-Pro, a multimodal reasoning LLM for protein function prediction, trained via SFT on synthetic reasoning traces and subsequent RL. I had a chance to interview @BoWang87 and @genophoria on their vision for the work and what comes next. Was fun to pick their brains on the bio! Check out the interview: youtube.com/watch?v=uZx3nU…

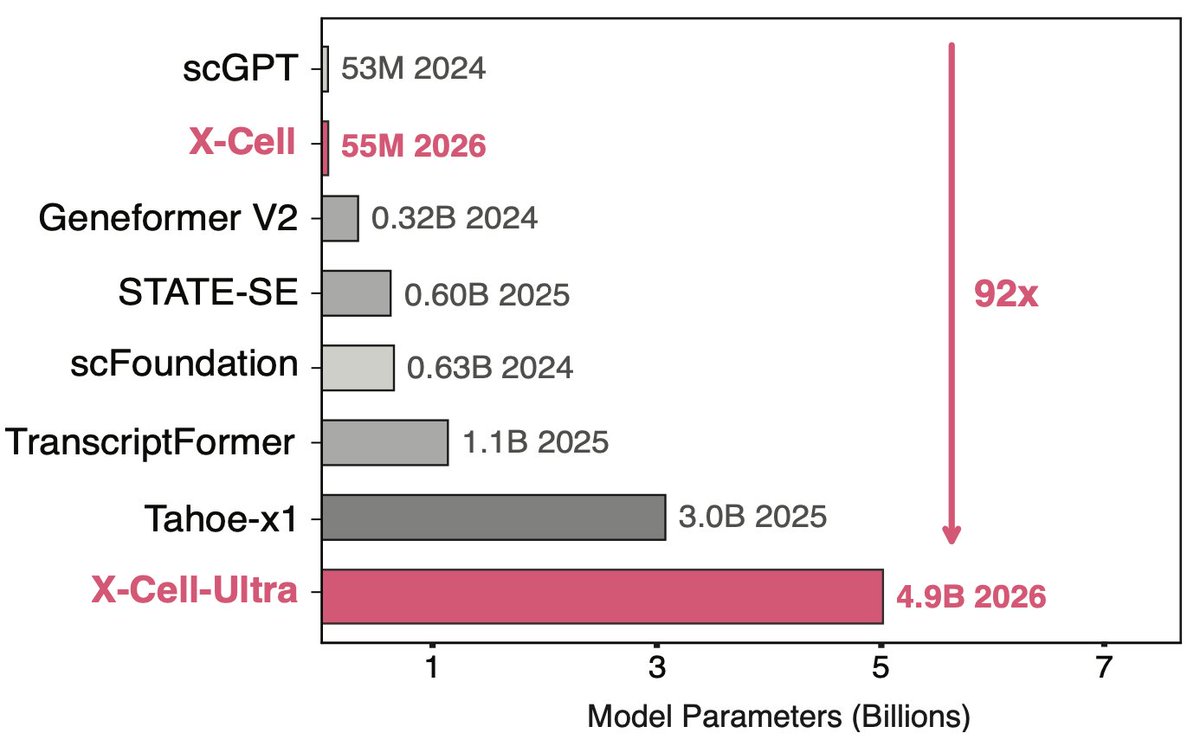

Our X-cell is up at @biorxiv_bioinfo ! Read our full paper at biorxiv.org/content/10.648… Part of the data and the model weights will be shared soon. stay tuned!



Today we’re announcing X-Cell — Xaira’s first step toward a virtual cell. 🧬 A foundation model that predicts how gene expression changes under causal perturbations — across cell types, conditions, and even unseen biology. This is not trained on observational atlases. It is trained on interventions. 🧵👇

Historic day for builders in bio: We’ve open-sourced @vevo_ai’s #Tahoe100M, largest single-cell atlas ever—by a wide margin—as the inaugural contribution to @arcinstitute’s Virtual Cell Atlas, ready for download today. A leap forward for AI models of cells & drug discovery. 🧵
