Neil Thomas
414 posts


Neil Thomas
@countablyfinite
Building AI for biological design @evoscaleai / @officialbiohub 🧬 Formerly: @Theteamatx 🌘 PhD @UCBerkeley 🙈


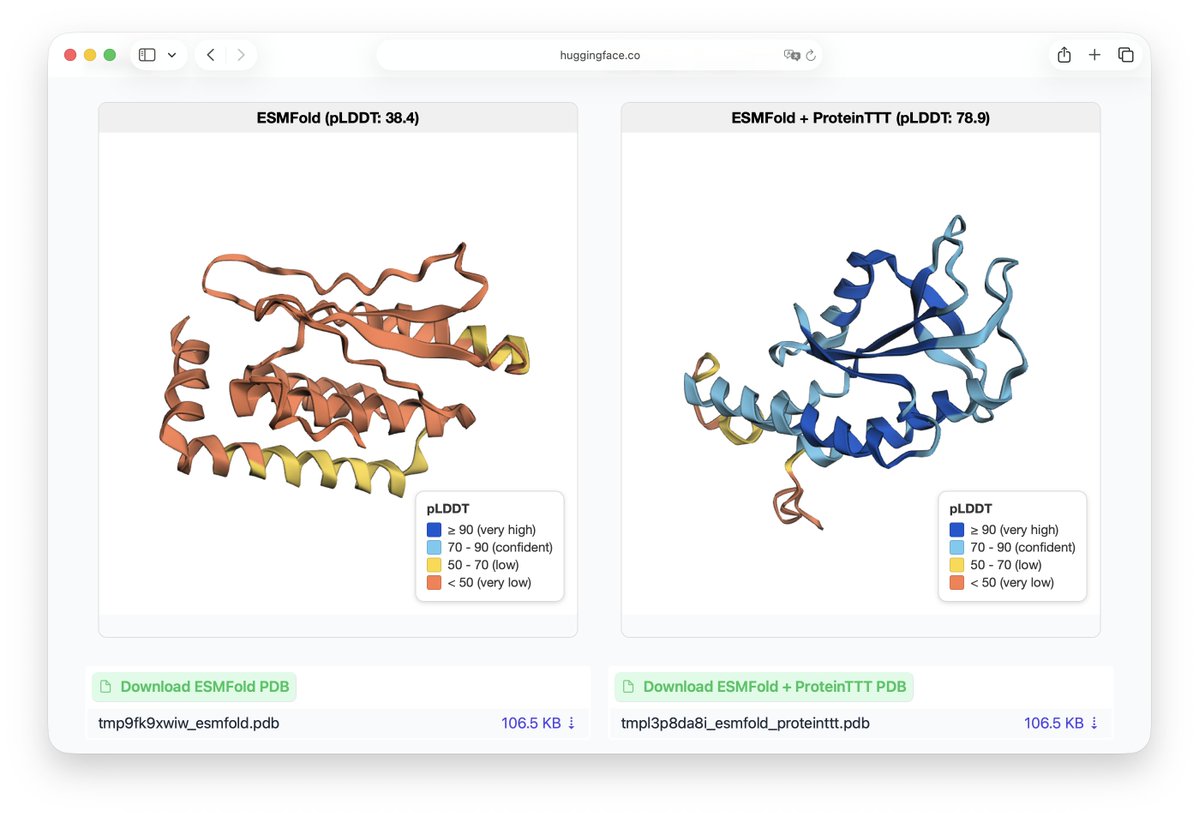





If you are a philanthropist, foundation or company and want to have a big impact on biological research and drug development, your biggest lever for impact is imo to fund labs creating large scale open data like @grocklin 's! Get your name on a dataset instead of a building! 😉

Hi Twitter 👋 We’re Evolved — organizers of the AI × Bio Hackathon (Nov 7–16). Today we’re announcing our judges: leaders from NVIDIA, OpenAI, FutureHouse, EvolutionaryScale, and more shaping the future of AI in biology.






1/4 🚀 Announcing the 2025 Protein Engineering Tournament. This year’s challenge: design PETase enzymes, which degrade the type of plastic in bottles. Can AI-guided protein design help solve the climate crisis? Let’s find out! ⬇️ #AIforBiology #ClimateTech #ProteinEngineering #OpenScience




Happy Sunday to all! This morning, we are excited to share Chase’s work developing a simple, scalable method to assemble 100s-1000s of custom genes from oligo pools using standard lab tools! (Small 🧵 below)