Biology+AI Daily@BiologyAIDaily
Sparse Autoencoders for Low-N Protein Function Prediction and Design
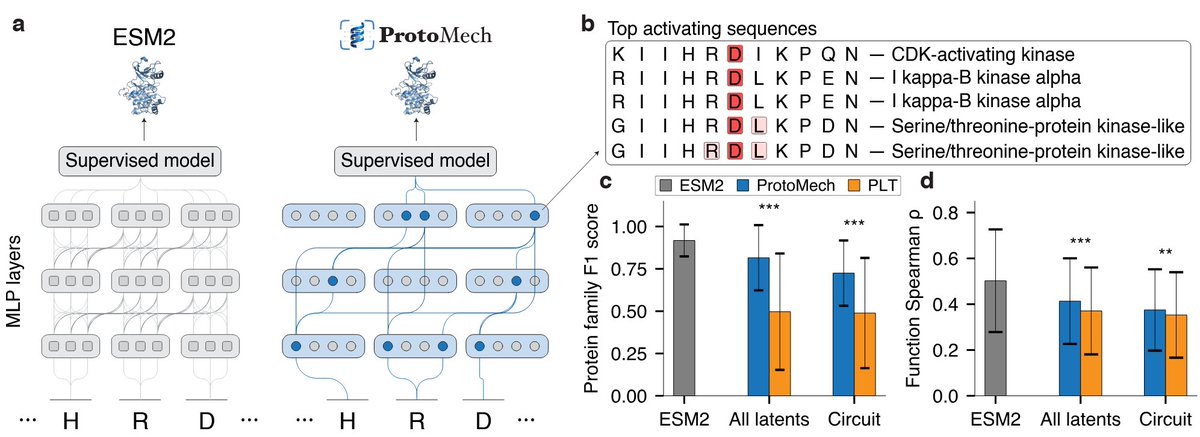
1. This study explores the use of Sparse Autoencoders (SAEs) for predicting protein function and designing proteins in low-data scenarios, demonstrating significant improvements over existing methods.
2. SAEs trained on fine-tuned ESM2 embeddings consistently outperform ESM2 baselines in fitness prediction tasks, even with as few as 24 sequences, showing their effectiveness in capturing biologically meaningful representations.
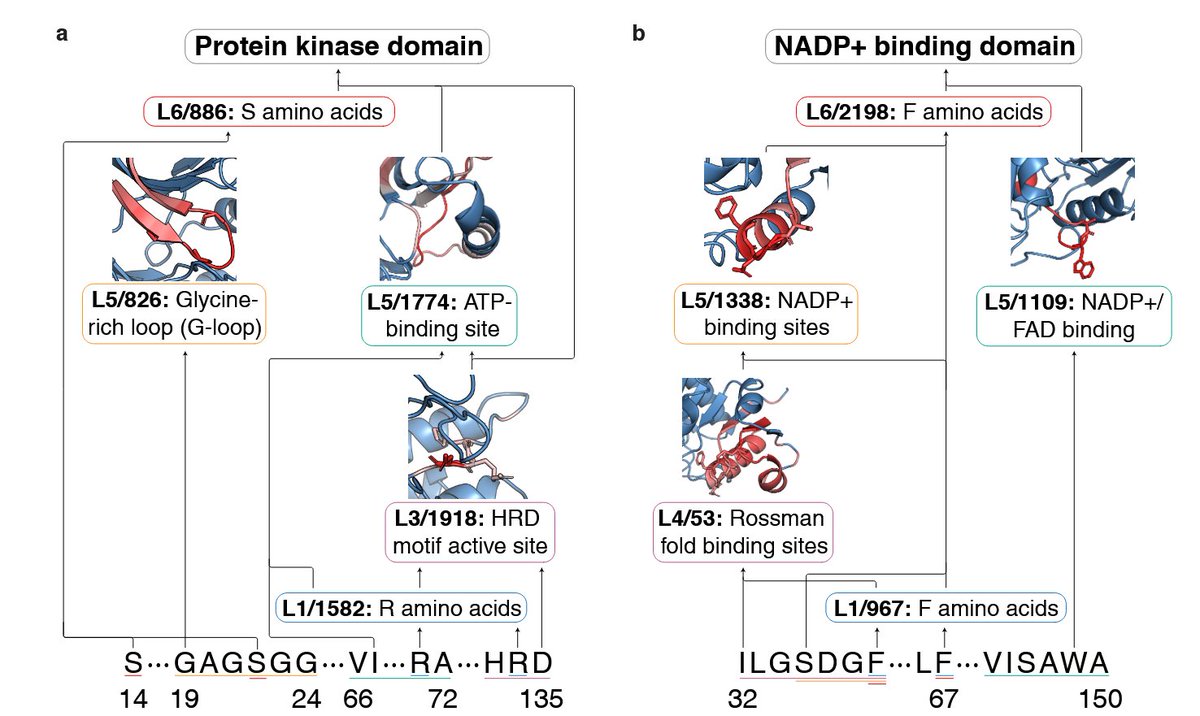
3. The study introduces a method to steer predictive latents in SAEs to design high-functioning protein variants, achieving top-fitness variants in 83% of cases compared to designing with ESM2 alone.
4. The authors analyze the best-performing variants in green fluorescent protein (GFP) and the IgG-binding domain of protein G (GB1), uncovering biologically meaningful motifs exploited by SAEs for steering.
5. SAEs achieve higher generalization to unseen variants compared to ESM2 in various low-N fitness extrapolation tasks, including position, regime, and score extrapolation.
6. The study highlights the importance of sparsity in SAEs, which compresses biologically relevant information into a sparse latent space, enhancing performance in low-N regimes.
7. The authors suggest future work could involve expanding the design space by steering multiple latents at once and coupling SAE steering with physics-based tools to optimize for both function and stability.
📜Paper: arxiv.org/abs/2508.18567…
💻Code: github.com/amirgroup-code…
#ProteinEngineering #MachineLearning #SparseAutoencoders #ProteinDesign #LowDataScenarios