Pablo Meyer
7.5K posts

Pablo Meyer
@jeriscience
Rabdomante @IBMresearch Escribí Genómica: https://t.co/e8g8y5DzAi… y Director @DR_E_A_M https://t.co/slIFTtQZsq https://t.co/8sKKshD8MD











BMFM-DNA: A SNP-aware DNA foundation model to capture variant effects 1. Researchers have developed BMFM-DNA, a groundbreaking approach to DNA language models that directly integrates Single Nucleotide Polymorphisms (SNPs) and other sequence variations during pre-training. This innovation addresses a key limitation of previous models that often overlooked the crucial biological impact of genomic variations. 2. The team pre-trained two Biomedical Foundation Models (BMFM) using ModernBERT: BMFM-DNA-REF, trained on reference genome sequences, and BMFM-DNA-SNP, which incorporates a novel representation scheme to encode sequence variations. This dual approach allowed for comprehensive evaluation of variation integration. 3. A significant innovation for BMFM-DNA-SNP involves mapping genetic variants from the dbSNP database, including SNPs, insertions, and deletions, to unique Chinese characters. This unique encoding implicitly introduces multiple nucleotide possibilities at a single genomic position, thereby expanding the effective pre-training DNA sample space and revealing hidden patterns of variation distribution. 4. Experiments showed that integrating sequence variations into these DNA language models leads to notable improvements across various fine-tuning tasks, demonstrating their enhanced ability to capture complex biological functions that are influenced by genomic differences. 5. The underlying architecture, ModernBERT, is a modernized encoder-only transformer model designed for improved efficiency and performance, particularly with longer sequence lengths. It incorporates features like Rotary Positional Embeddings (RoPE) and FlashAttention. 6. To support further research and community contributions, the models and the code for reproducing the results have been publicly released through HuggingFace and GitHub. A comprehensive software package, bmfm-multi-omic, is also available for pre-training, finetuning, and benchmarking genomic foundation models. 💻Code: github.com/BiomedSciAI/bi… 📜Paper: arxiv.org/pdf/2507.05265… #ComputationalBiology #Genomics #FoundationModels #SNPs #DNAModels #Bioinformatics #AI #MachineLearning




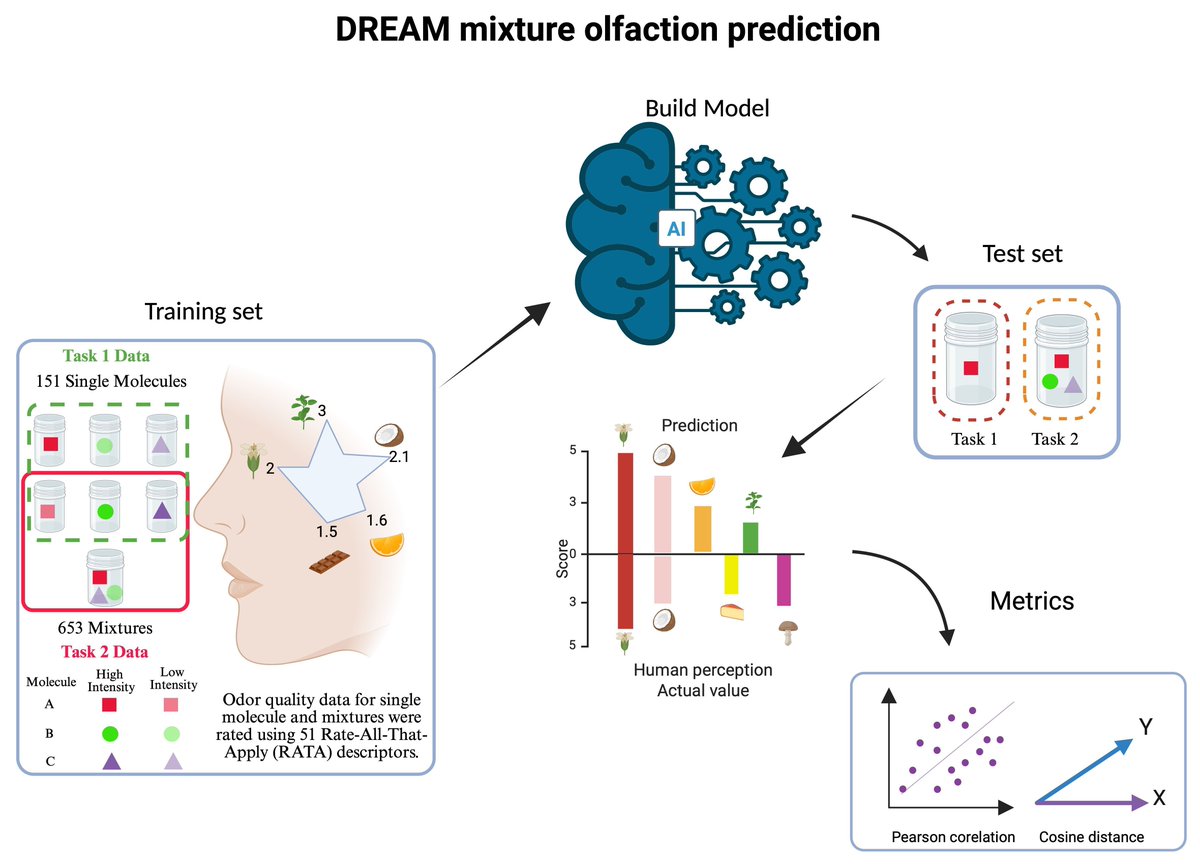
With a contemporary smartphone, capturing, transmitting, and replicating sounds and visuals is a breeze, yet replicating scents remains elusive. If you want to change this, consider joining the 2025 @DR_E_A_M Olfaction Challenge at synapse.org/Synapse:syn647…. @jeriscience