Jeff Wintersinger
1.8K posts


Jeff Wintersinger
@jwintersinger
Senior Research Scientist in comp bio at @deepgenomics. Did PhD work on cancer evolution. My three favourite things are 🍌s, riding my 🚴fast, and vim.







2025 Machine Learning in Computational Biology (#MLCB) meeting starts TODAY (9/10) at 9:30 (EST)! We have a great lineup of keynotes, contributed talks, and posters today and tomorrow! Schedule: mlcb.org/schedule Join for free via livestream: @mlcbconf" target="_blank" rel="nofollow noopener">m.youtube.com/@mlcbconf




KV caching overcomes statelessness in a very meaningful sense and provides a very nice mechanism for introspection (specifically of computations at earlier token positions) the Value representations can encode information from residual streams of past positions without significant compression bottlenecks before they're added to residual streams of future positions the greatest constraint here imo is that it doesn't provide longer *sequential* computational paths that route through previous states, but it does provide a vast number of parallel computational paths that carry high dimensional (proportional to the model's hidden dimension) stored representations from all earlier layers/positions yes, some of the information in intermediate computations e.g. in the MLP is compressed and cannot be reconstructed fully, but that's just how any reasonable brain works if accurate introspection of previous states is incentivized at all, you should expect this mechanism to be exploited for that. and I think it definitely is, like, being able to accurately model your past beliefs and intentions and articulate them truthfully is pretty fucking useful for coordinating with yourself across time and doing useful cognitive work over multiple timesteps; hell, it's useful for writing fucking rhyming poems. also if you have interacted with models you may observe empirically that introspective reporting yields remarkably consistent results, and this is more true of more capable models with skillful agentic posttraining, which are necessarily minds that intimately know the shape of themselves in motion.



Blank Bio (@blankbio_) is building foundation models to power a computational toolkit for RNA therapeutics, starting with mRNA design and expanding to target ID, biomarker discovery, and more. ycombinator.com/launches/O8Z-b… Congrats on the launch, @hsu_jonny, @phil_fradkin & @ianshi3!


Today, I'm excited to present the second big chunk of my Ph.D. work. We are building a thorough assessment of the burgeoning field of gene expression forecasting. Something is deeply wrong.🧵 - Preprint: biorxiv.org/content/10.110… - Code: #a-systematic-comparison-of-computational-methods-for-expression-forecasting-with-pereggrn" target="_blank" rel="nofollow noopener">github.com/ekernf01/pertu…


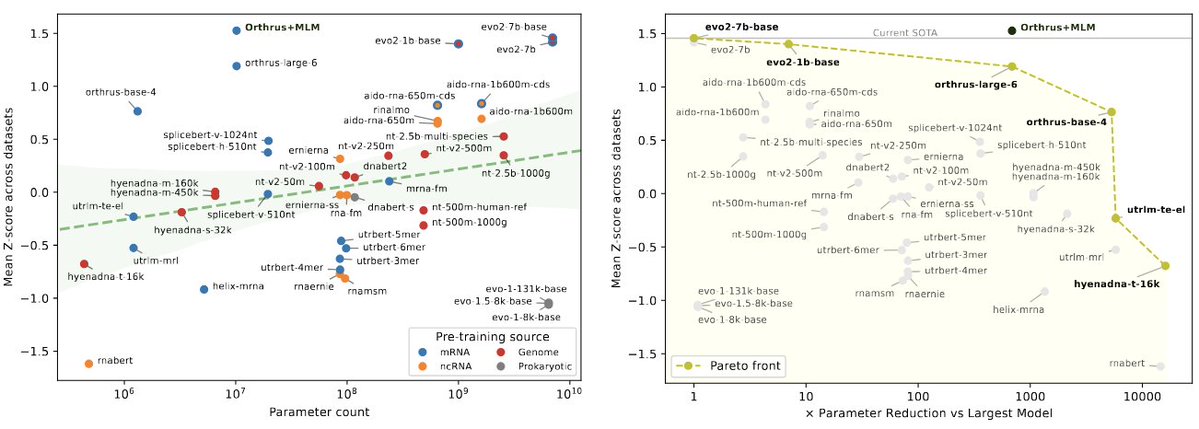
We're excited to release 𝐦𝐑𝐍𝐀𝐁𝐞𝐧𝐜𝐡, a new benchmark suite for mRNA biology containing 10 diverse datasets with 59 prediction tasks, evaluating 18 foundation model families. Paper: biorxiv.org/content/10.110… GitHub: github.com/morrislab/mRNA… Blog: blank.bio/post/mrnabench