Lamin retweetledi
Lamin
13 posts

Lamin
@laminlabs
Open data framework for biology. Context and memory for datasets and models at scale.
Katılım Aralık 2021
6 Takip Edilen200 Takipçiler
Lamin retweetledi

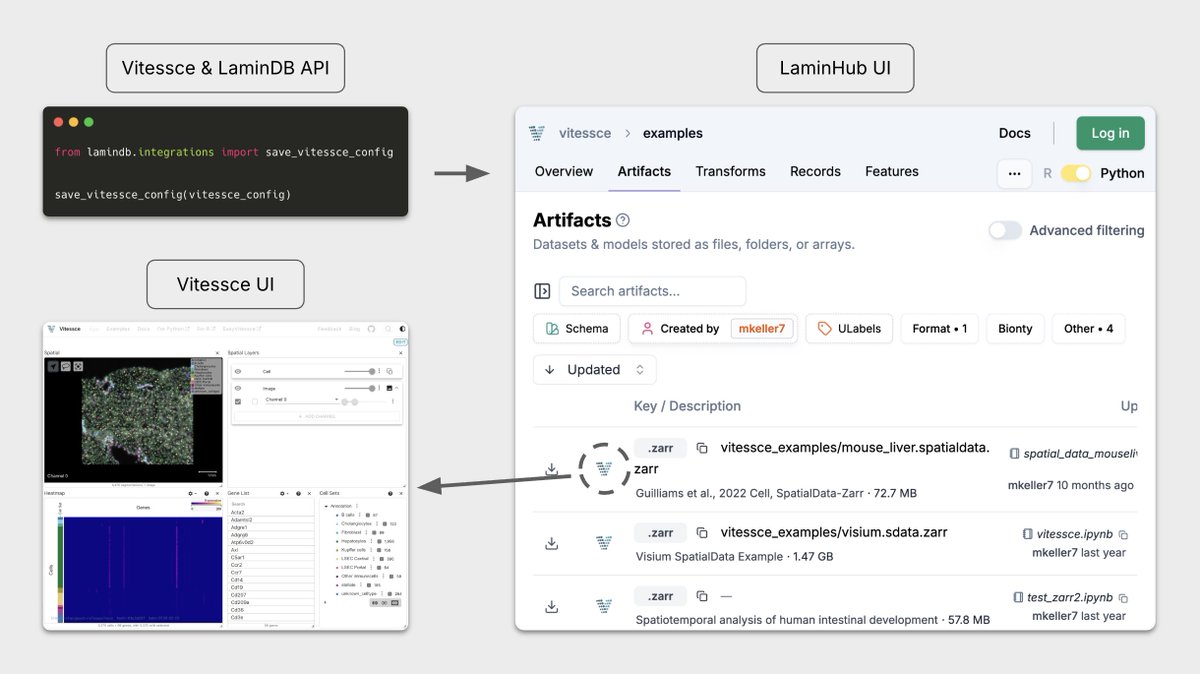
Two years ago we partnered with Mark Keller from Nils Gehlenborg’s Lab at Harvard to make Vitessce work seamlessly with LaminDB for interactive visualization of multimodal + spatial datasets.
The integration has found much use across academia, biotech, and pharma — so we wrote up on design principles & use cases.
This was a team effort involving Altana, Richard & Sunny in addition to Mark.
Read the post: blog.lamin.ai/vitessce
English
Lamin retweetledi
Lamin retweetledi

Nice, detailed benchmark of backends that allow for batched training on a large scRNA-seq corpus - efficiently dealing with the specifics of a scenario can be a big engineering challenge, lowering this barrier will enable cool computational biology down the road!
Alex Wolf@falexwolf
What's a good way of organizing scRNA-seq data for training foundation models? Say you run 1k experiments and each measures counts for 1M cells with varying metadata and orthogonal data. Storing these data in one gigantic array isn’t exactly easy. We wondered whether it’s necessary to train foundation models and found 3 setups that made sense to us. lamin.ai/blog/arrayload…
English
Lamin retweetledi

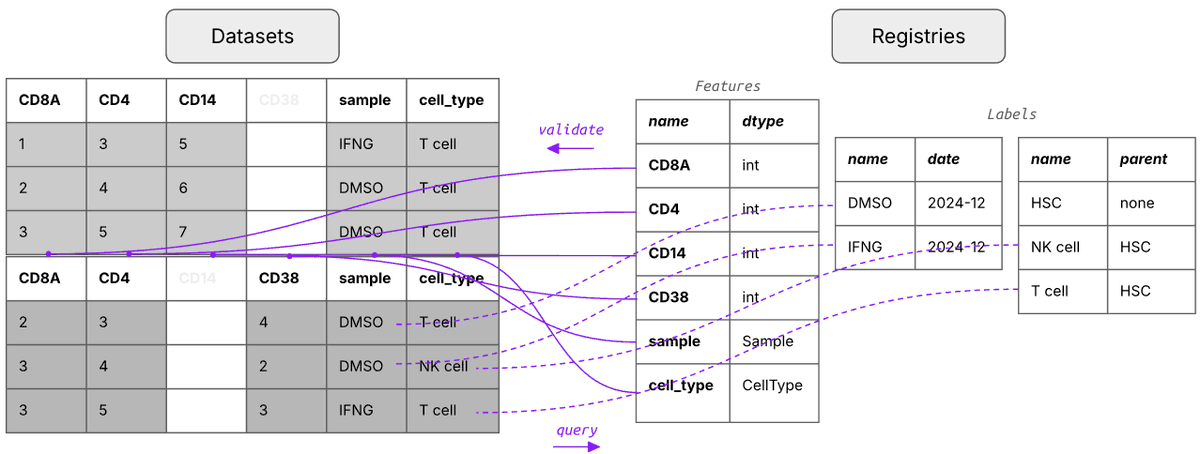
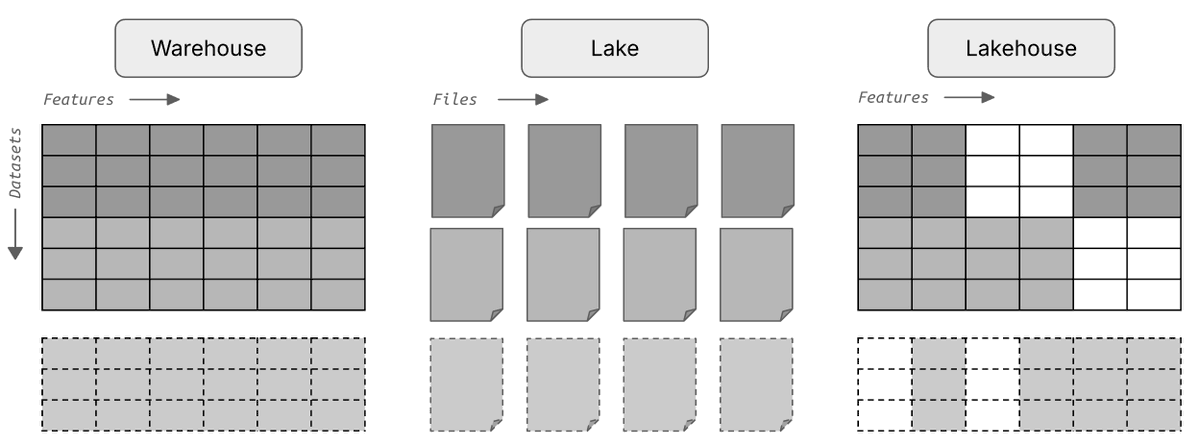
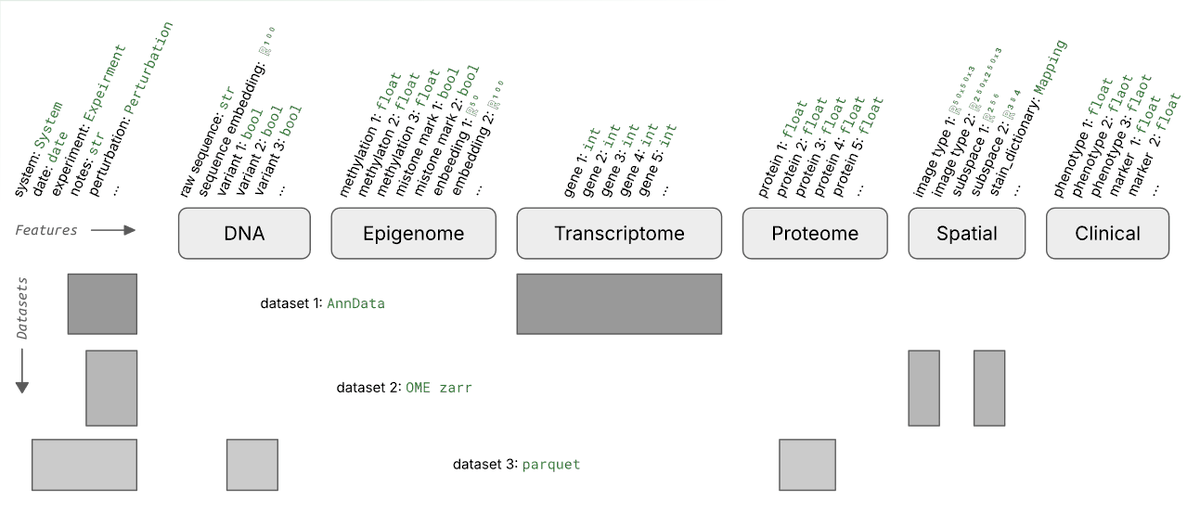
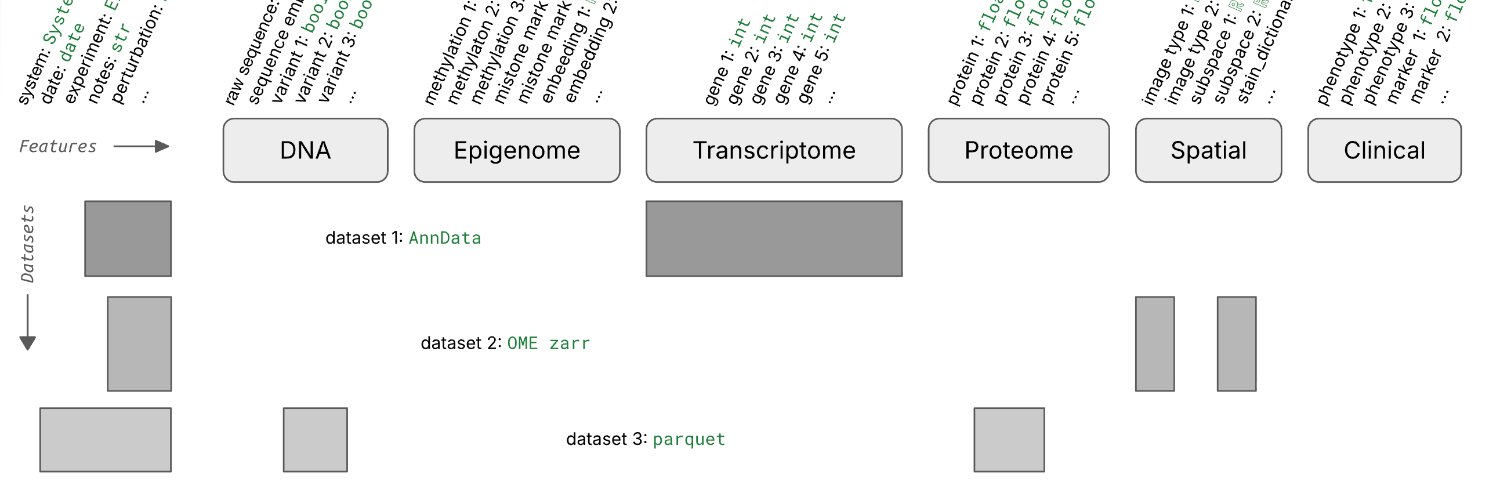
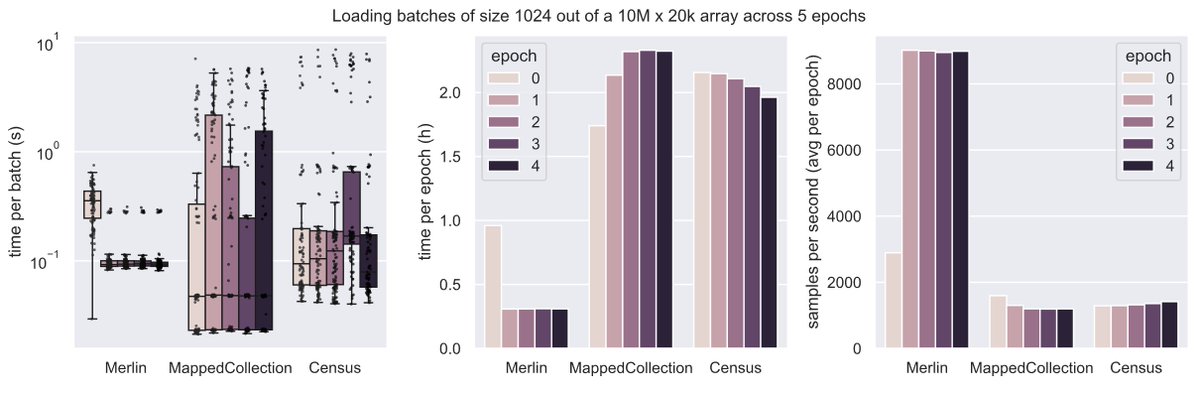
What's a good way of organizing scRNA-seq data for training foundation models?
Say you run 1k experiments and each measures counts for 1M cells with varying metadata and orthogonal data.
Storing these data in one gigantic array isn’t exactly easy.
We wondered whether it’s necessary to train foundation models and found 3 setups that made sense to us.
lamin.ai/blog/arrayload…
English
Lamin retweetledi

Organizing scRNA-seq data with LaminDB - nxn.se/valent/2024/1/…
Indonesia
Lamin retweetledi

Thank you for the awesome collaboration, @marenbuettner!
With Pytometry, we'd like to share readfcs: A package to load data and metadata from FCS files to AnnData.
pip install readfcs
English
Lamin retweetledi