Sabitlenmiş Tweet
Laure Ciernik
30 posts

Laure Ciernik
@lciernik
PhD student at ML Group @TUBerlin @bifoldberlin @HFA_academy @ELLISforEurope | MSc thesis at Boeva Lab, @ETH_en
Katılım Eylül 2023
149 Takip Edilen90 Takipçiler

Large thanks for the vital support: @bifoldberlin, @HFA_academy, @uni_tue, Tübingen AI Center, @HelmholtzMunich, @MunichCenterML, and @AIgnostics. We couldn't have done it without this ecosystem🚀
English

7/ Huge thanks to my shared first author, Marco Morik, and all great co-authors:
@lukas_thede, @LucaEyring, Shinichi Nakajima, @zeynepakata, and @lukas_mut 🙏
English

🎉 Update: This work got accepted to #icml2025!!
Huge thanks to my amazing co-authors @LorenzLinhardt, Marco Morik, @jdppel, @skornblith, and @lukas_mut for their great work and to all collaborators! 🙏
📄 Paper: arxiv.org/abs/2411.05561
💻 Code: github.com/lciernik/simil…
🧵1/3

Laure Ciernik@lciernik
If two models are more similar to each other than a third on ImageNet, will this hold for medical/ satellite images? Our preprint analyzes how vision model similarities generalize across datasets, the factors that influence them, and their link to downstream task behavior. 🧵1/7
English
Laure Ciernik retweetledi

1/🚀 Excited to share RegVelo, our new computational model combining RNA velocity with gene regulatory network (GRN) dynamics to model cellular changes and predict in silico perturbations. Here's how it works and why it matters! 🧵👇
biorxiv.org/content/10.110…

English
Laure Ciernik retweetledi

🚀 New preprint from our lab, @ekrym2 and @fabian_theis : UniversalEPI, an attention-based method to predict enhancer-promoter interactions from DNA sequence and ATAC-seq🌟
🔬 Key Highlights:
- Predicts chromatin interactions across unseen cell types with no retraining.
- Outperforms state-of-the-art models like C.Origami and ChromaFold with Spearman’s Rho > 0.92 in unseen cell types.
- Tracks chromatin dynamics in processes like macrophage reprogramming and cancer transcriptional states.
- Scalable for both bulk and single-cell chromatin accessibility data.
🧬UseUniversalEPI for in-silico 3D chromatin modeling in your favorite model! Examples of applications: studies on genetic diseases, cell differentiation, and the regulatory impacts of non-coding variants.
📄 Read the full preprint: doi.org/10.1101/2024.1… by @AayushGrover8 @ekrym2 @ilibarra @fabian_theis et al.

English
Laure Ciernik retweetledi

1/ 🧬 Single-cell genomics reveals biological variations beyond cell types. Unveiling these in separate latent dimensions is known as disentanglement. Led by @AmirAliMoinfar, we introduce DRVI to learn nonlinear, disentangled & interpretable latent spaces.
biorxiv.org/content/10.110…
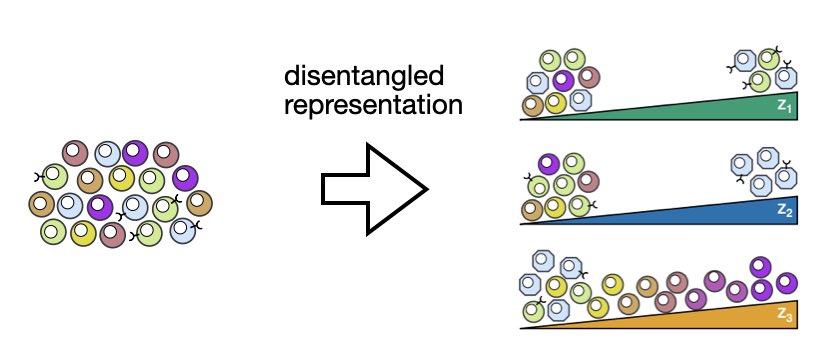
English
Laure Ciernik retweetledi

❓ Do histopathological foundation models eliminate batch effects? ❓
The surprisingly clear answer is: they do not!
Find out more in our new paper that we will present at the AIM-FM Workshop at @NeurIPSConf.
Link: arxiv.org/abs/2411.05489

English

Want to learn more? Read our new preprint:arxiv.org/abs/2411.05561 ✨
Deeply grateful to @LorenzLinhardt, Marco Morik, @jdppel, @skornblith, and @lukas_mut for their rigorous work and dedication throughout this project. Each contribution was essential to make this possible! 🙏 7/7
English
