Luke Funk
218 posts


Luke Funk
@lukebfunk
@CajalNeuro. Former: @BlaineyLab, @broadinstitute, @MIT_HST. Opinions my own. https://t.co/aDN2eJDKCK
Seattle, WA Katılım Eylül 2018
426 Takip Edilen185 Takipçiler
Luke Funk retweetledi

Excited to share how @jacobborrajo Anna Le and I are enabling the future of profiling dynamic cellular processes! We built a synthetic RNA export pathway for cells to "self-report" their transcriptional states in real-time. Big thanks to the entire team!
biorxiv.org/content/10.110…
GIF
English

@markkitti @manorlaboratory I'm here. Tried to use h5 for a while, but I think zarr is the future.
English

Closest I have found to what I am looking for is MolArt (cusbg.github.io/MolArt/), but doesn't seem to allow custom annotations.
English

@mike_tilapia @DrAnneCarpenter Awesome work! I might be missing it, but I am not seeing the supplementary table on bioRxiv...is it posted somewhere?
English

Our latest preprint is out! In a fun collaboration with @DrAnneCarpenter, we conducted a systematic high-content screen to understand the role of protein mislocalization in diverse human disorders. 1/8
biorxiv.org/content/10.110…
English

I'm really excited to present work from @ophirshalem’s group with my outstanding co-first author Yevgeniy Serebrenik @YSereb to tag and manipulate proteins at scale, which we use to study the #proteome and #proteostasis biorxiv.org/content/10.110…
English

@stephsansbury @ophirshalem @YSereb Congrats on getting this out @stephsansbury; excited to read in detail!
English
Luke Funk retweetledi

How can machine learning help you to analyze your microscopy images?
Find out at the "Deep Learning for Microscopy Image Analysis" course at the @MBLScience in Woods Hole from Aug 21-Sep 5!
Applications are due 🚨 May 18 🚨
Read on for more! 👇
mbl.edu/education/adva…
English

I am so excited to share that I will be starting a lab in January 2024 as an Assistant Professor at @OHSUSOM in Portland! I am building my team, so if you are interested in cellular cryoelectron tomography, computational structural biology, and working in PDX, hit me up!

English
Luke Funk retweetledi
Luke Funk retweetledi

@jonesr18 @nika_shakiba I see, thanks for the clarification! Are the supplementary figures available?
English

@nika_shakiba @lukebfunk The v4 one already caused reduced growth rates - if Oct4 levels continue to go up, the efficiency would likely decrease due to this!
English

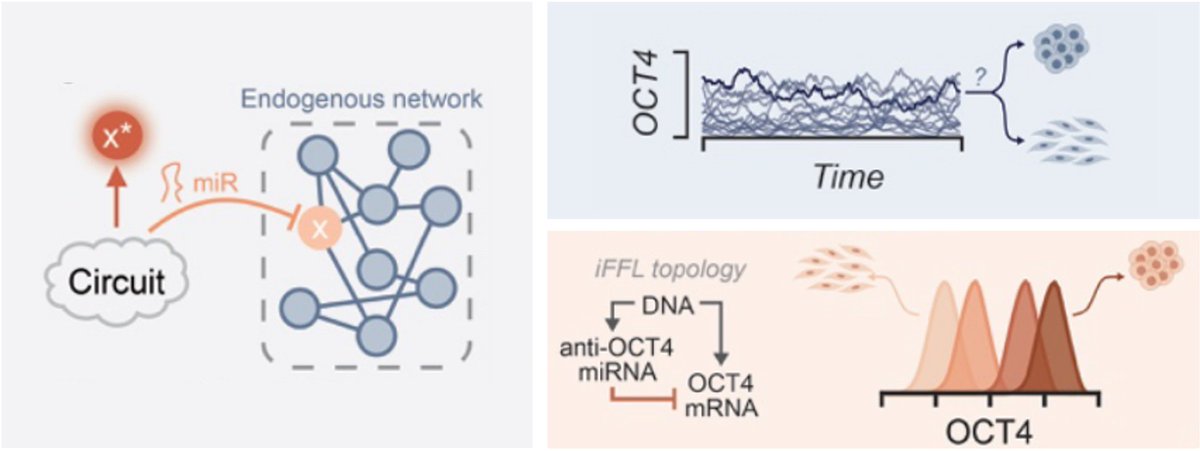
🚨 Check out our new preprint!
Super excited to see this go over the finish line, thanks to a collaborative effort 📢 Katherine & Trevor!
Tl;dr: #SynBio genetic circuits 🧬 + reprogramming to #iPSCs 🧫= robust cell fate control
See thread for the story!
biorxiv.org/content/10.110…
English

@nika_shakiba Also, in the experiments in Fig. 4c & d, did you compare to a control without the miR casette? Wondering if the v4 miR-Oct4e with many mismatches is actually producing any repression? Or if the high & tight Oct4 distribution is driven primarily by the high MOI?
English

@nika_shakiba In the final trajectory controller designs in Fig. 4a, it looks like the miR-Oct4e variants will target both the endogenous and exogenous OCT4 mRNA? Did you have a way to separately control and “overwrite” Oct4 levels in these experiments like earlier in the paper?
English

@RecursionChris Also, how were the labeled genes chosen? Random subset? Intentionally selected for specific properties? What properties?
English

@RecursionChris Totally agreed! Just not sure what to think of the 16k anonymized genes without any specific plans to de-anonymize in the future.
English

Can't decide if this is a useful dataset release for external users
Chris Gibson@RecursionChris
Big day for open-science at Recursion! Releasing a 2.2M image, 100TB genome-spanning CRISPR KO dataset* in primary human cells alongside a demo of one of our internal apps. Kudos to the team for getting this ready! Can’t wait to see what the community finds/builds on top of it!
English




