M⌬nica retweetledi

Japan team discovers why cats leave meals unfinished japantimes.co.jp/news/2026/04/0…
English
M⌬nica
356 posts


@mon_difer
A universe of atoms, an atom in the universe. #Postdoc #PhD👩🏻🔬 #Science #NMR #Spectroscopy 📈📉 #OrganicChemistry 🧪#NaturalProducts 🌿












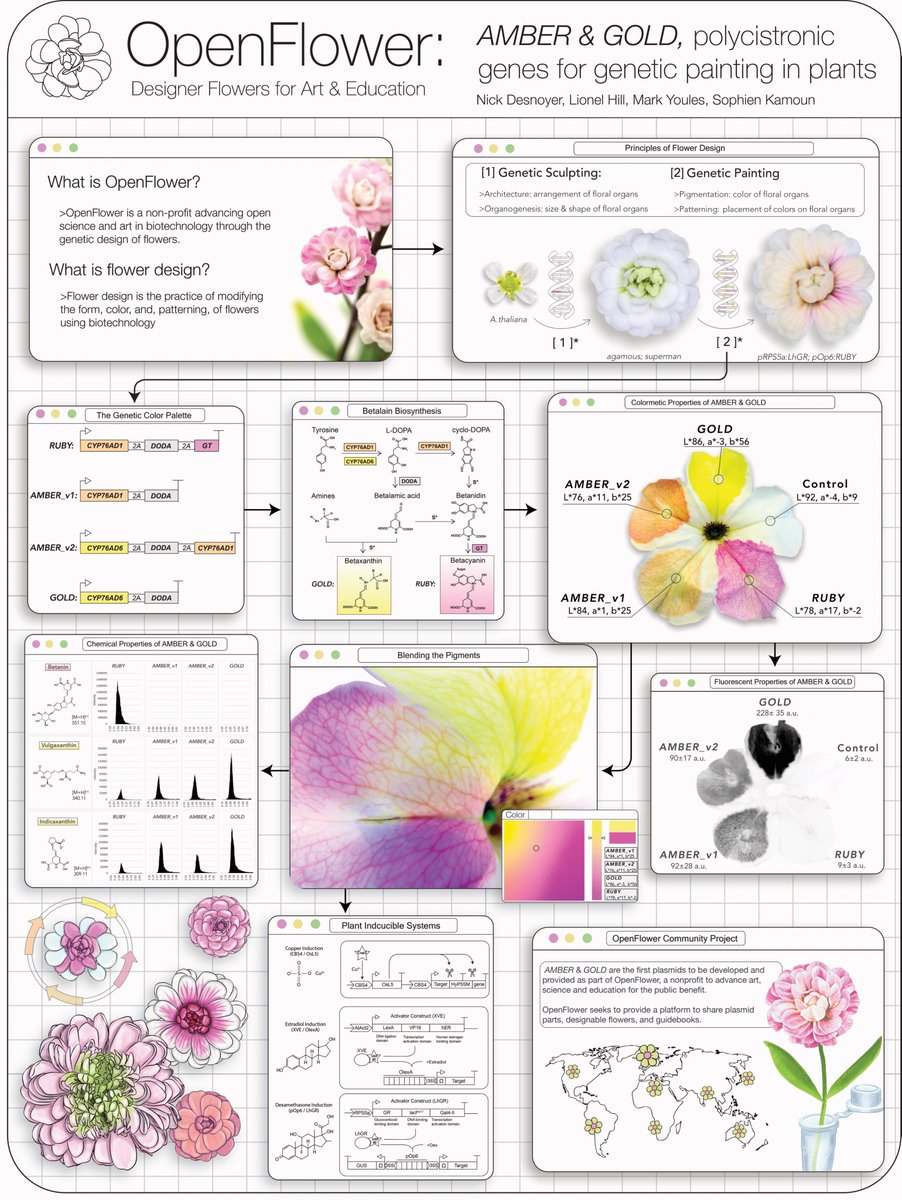
My poster on flower bioengineering at the #AIEBaB26 conference in Bristol last week 🌹 🧬 ✨