Sabitlenmiş Tweet

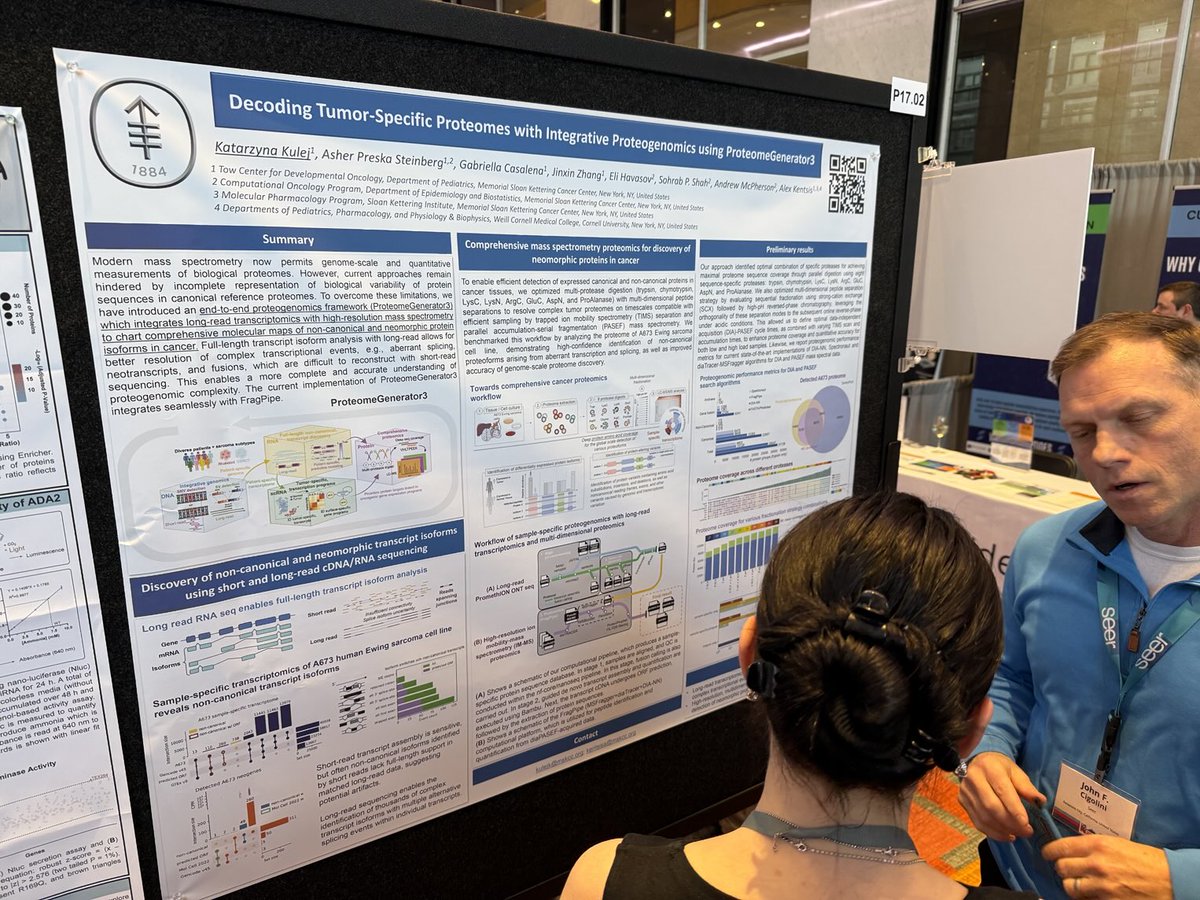
Friends, #FragPipe 22 has been released, and it's a big update! diaTracer enables spectrum-centric analysis of diaPASEF data. Skyline integration. Koina server for more deep-learning prediction options. DDA+ mode for ddaPASEF. DIA glycoproteomics and new chemoproteomics workflows

English