Introducing ProtSpace: a tool for visualizing protein embeddings in 2D/3D! 🎨
biorxiv.org/content/10.110…
Try it:
📦github.com/tsenoner/prots…
🌐protspace.rostlab.org
#Proteomics #DataViz
English
RostLab
499 posts


@rostlab
Predict protein structure and function using sequence, evolution & more! Operated by lab members.


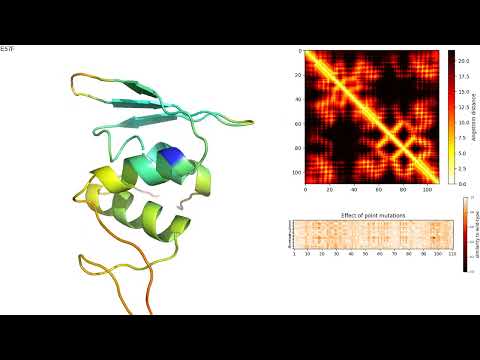
VespaG: Expert-guided protein Language Models enable accurate and blazingly fast fitness prediction biorxiv.org/cgi/content/sh… #biorxiv_bioinfo



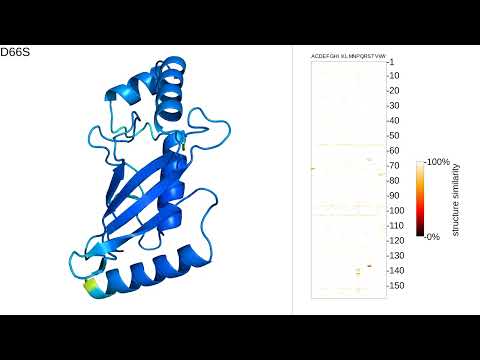




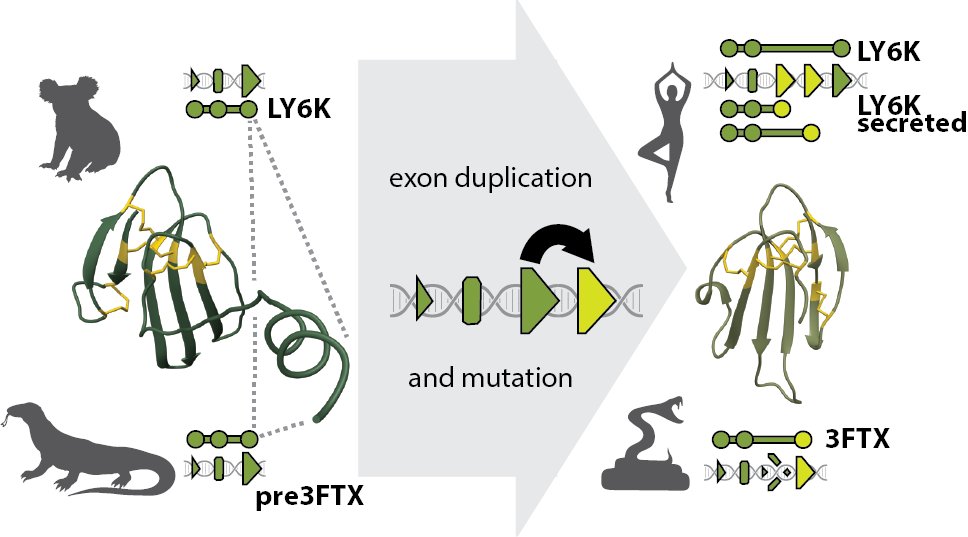






Alignment-based protein mutational landscape prediction: doing more with less biorxiv.org/cgi/content/sh… #bioRxiv