Tom Barrett retweetledi

🚀 Introducing Nucleotide Transformer v3 (NTv3)
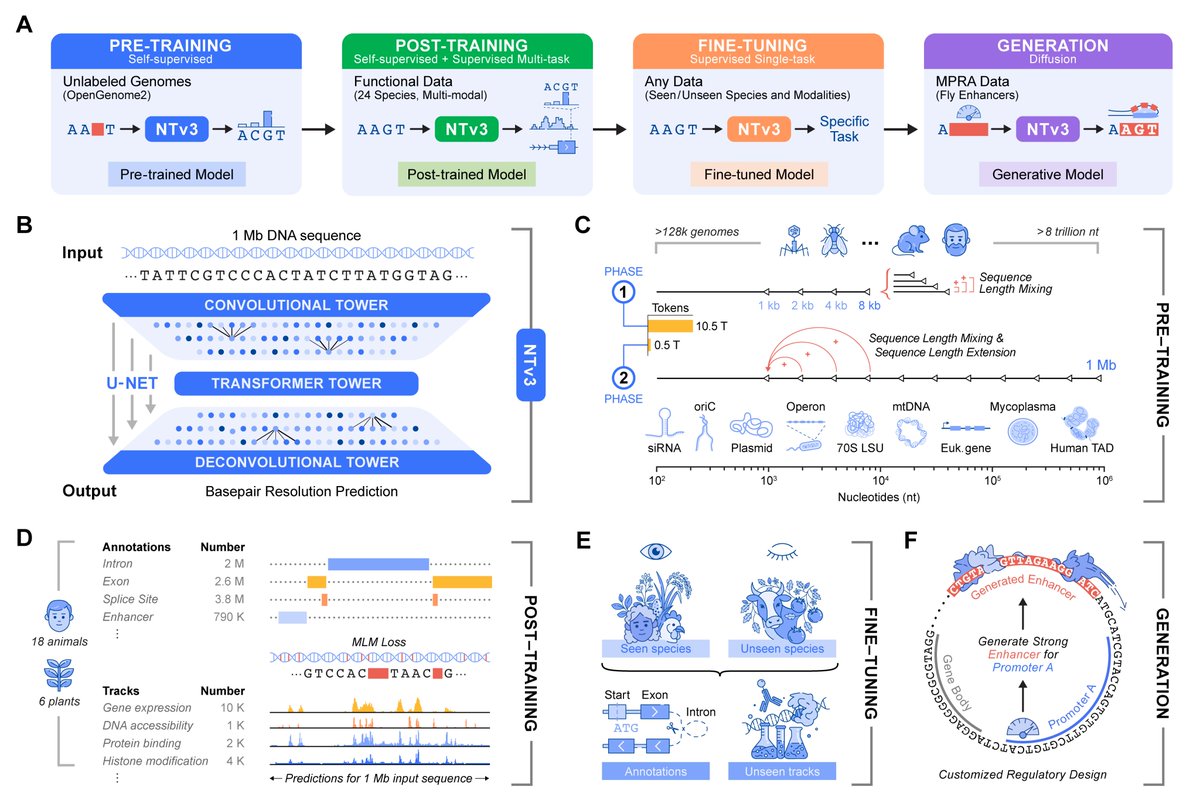
Today, we are very excited to share our latest foundation model for biology - Nucleotide Transformer v3 (NTv3).
NTv3 is @instadeepai new multi-species genomics foundation model, designed for 1 Mb, single-nucleotide-resolution prediction, and for bridging representation learning, sequence-to-function modeling, and generative regulatory design within a single framework 🧬
This work was developed in close collaboration with @AlexanderStark8 , @volokuleshov , and @pkoo562 , and reflects several years of joint effort at the intersection of machine learning and regulatory genomics.
English












