Will Connell
310 posts

Will Connell
@wilstc
predicting phenotypes 🖥🧬🔮 @transcriptabio

Manufacturing-aware generative models enable petascale synthesis of designed DNA go.nature.com/3NxXt1I

We mapped gene interactions across different environmental conditions (GxGxE) at scale for the first time in human cells. These maps lead to the realization that many genes function in a context dependent manner which provides insight into how humans have relatively few genes but many cell types. Congratulations Ben! Paper: cell.com/molecular-cell…









I will not stop tweeting this until every drug discovery company gets human evidence in 3 years or less

R&D productivity for new drug approvals has steadily declined over the past 50 years. With the rise of AI tools, a common belief is that they will dramatically boost research productivity by accelerating the "search" for solutions—a sentiment echoed widely at #JPM this week. 🧵







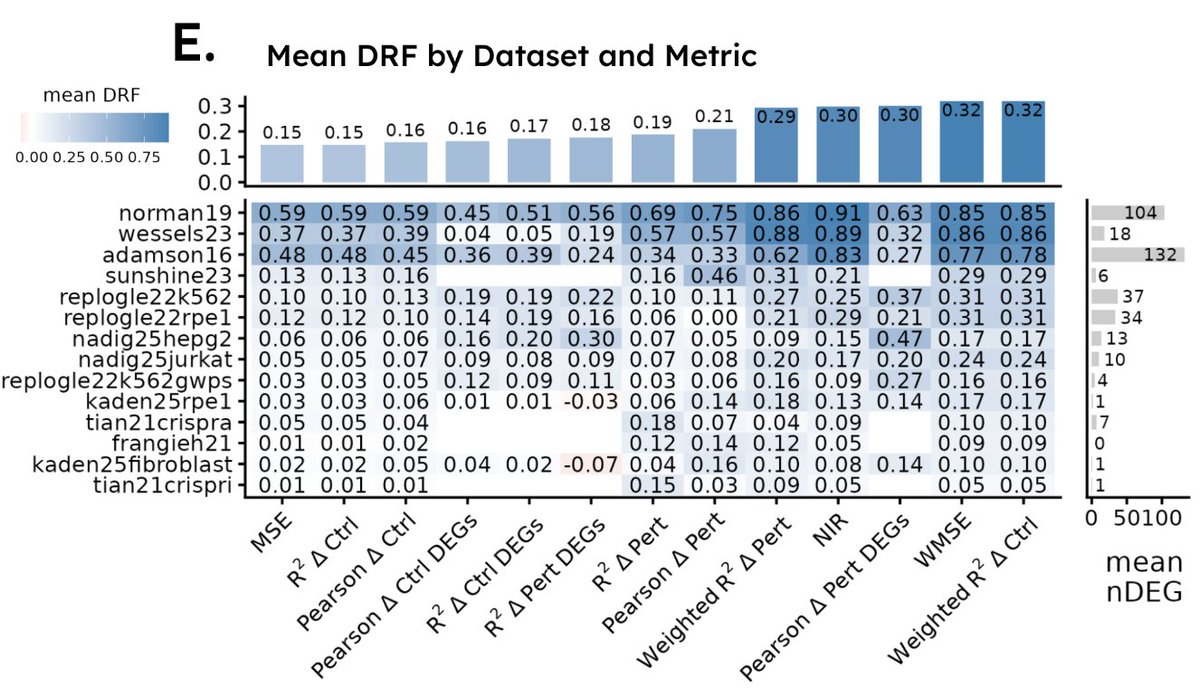


My second post on the Arc Virtual Cell Challenge. The challenge’s Discord forums are in turmoil. Some participants have discovered a trick to get to the top of the leaderboard. gmdbioinformatics.substack.com/p/arc-virtual-… #arc_virtual_cell_challenge #foundation_models




Virtual Cell community - this one's for you! X-Atlas/Orion is now live on Hugging Face. Train your own models with streamlined workflows built into the Hugging Face API. 🔗 HuggingFace: huggingface.co/datasets/Xaira… 📜 License: cc-by-nc-sa-4.0

Welcome to the age of generative genome design! In 1977, Sanger et al. sequenced the first genome—of phage ΦX174. Today, led by @samuelhking, we report the first AI-generated genomes. Using ΦX174 as a template, we made novel, high-fitness phages with genome language models. 🧵

🌎👩🔬 For 15+ years biology has accumulated petabytes (million gigabytes) of🧬DNA sequencing data🧬 from the far reaches of our planet.🦠🍄🌵 Logan now democratizes efficient access to the world’s most comprehensive genetics dataset. Free and open. doi.org/10.1101/2024.0…


