Che,Yang
85 posts




📣 We're excited to announce the opening of multiple faculty positions! Do you use and/or develop artificial intelligence methods and mechanistic mathematical models to address fundamental questions in biology? We'd love to hear from you. (Please share far and wide 🙏)


Exploring "dark matter" protein folds using deep learning biorxiv.org/cgi/content/sh… #biorxiv_bioinfo
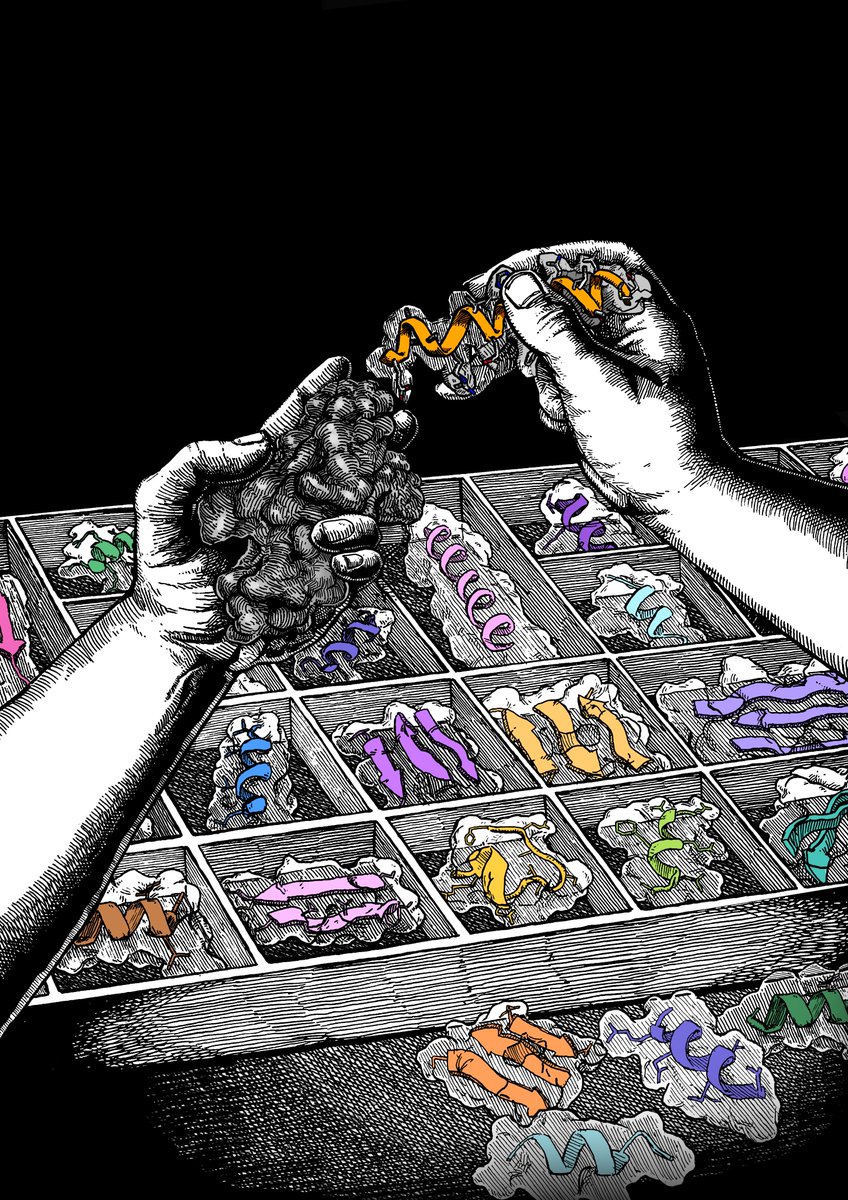
















Using local geometries and a simple string encoding to generate existing and novel protein folds. Really happy this work w/ @zanderharteveld, @yangche7, @SesterhennF, Stéphane and @befcorreia is finally out! pnas.org/doi/10.1073/pn…


Our last explorations with MaSIF and surface fingerprints for the computational design of de novo protein-protein interactions ! A lot of hard work from Pablo, @AKVanHall, Sarah, Andrea, Anthony, @mmbronstein and collaborators biorxiv.org/content/10.110…


Huge congrats to our dear Bruno @befcorreia Correia for achieving tenure at the @EPFL_en: as of this October 1, he is Associate Professor of Bioengineering! Lucky us at @EPFL_BioE: now we can look forward to many more years with such a gem of a Colleague! actu.epfl.ch/news/bruno-cor…

We’ve been having a lot of fun with these switches! Thank you guys #lpdi for so much hard work specially Sailan @Scheller123 @yangche7 Pablo and good collaborators @saireddy911


Huge congrats to our dear Bruno @befcorreia Correia for achieving tenure at the @EPFL_en: as of this October 1, he is Associate Professor of Bioengineering! Lucky us at @EPFL_BioE: now we can look forward to many more years with such a gem of a Colleague! actu.epfl.ch/news/bruno-cor…