Yu Zhao
129 posts

Yu Zhao
@yu48693
Postdoctoral Research Fellow, Bioinformatics, Systems Immunology @BrighamWomens @broadinstitute

@michecurtis and I are thrilled to share our preprint "Reproducible single cell annotation of programs underlying T-cell subsets, activation states, and functions" from the @soumya_boston lab @BrighamResearch and @broadinstitute! biorxiv.org/content/10.110… 1/12





Early and late RNA eQTL are driven by different genetic mechanisms biorxiv.org/content/10.110… #biorxiv_genomic





We are excited to share Genotype-Neighborhood Associations, GeNA, a new tool adapting our CNA framework to detect cell states associated in abundance with genetic variants at genome-wide scale in high-dimensional single-cell data w/ @soumya_boston doi.org/10.1101/2023.1… 🧵

We are happy to announce that our new preprint regarding the scLinaX, software to quantify escape from X chromosome inactivation from 10X scRNA-seq data, was released from bioRxiv 🎉🎉🎉!! biorxiv.org/content/10.110… (1/N)



So excited to share our #RheumatoidArthritis tissue chromatin atlas of 87K cells from 30 individuals! With, @soumya_boston, @saorisakaue, @aparnanathan, @Donlinlab, Kevin Wei and #AMP-RA! @NIH_NIAMS. biorxiv.org/content/10.110… (Tweeting on behalf of the talented Kathryn Weinand)

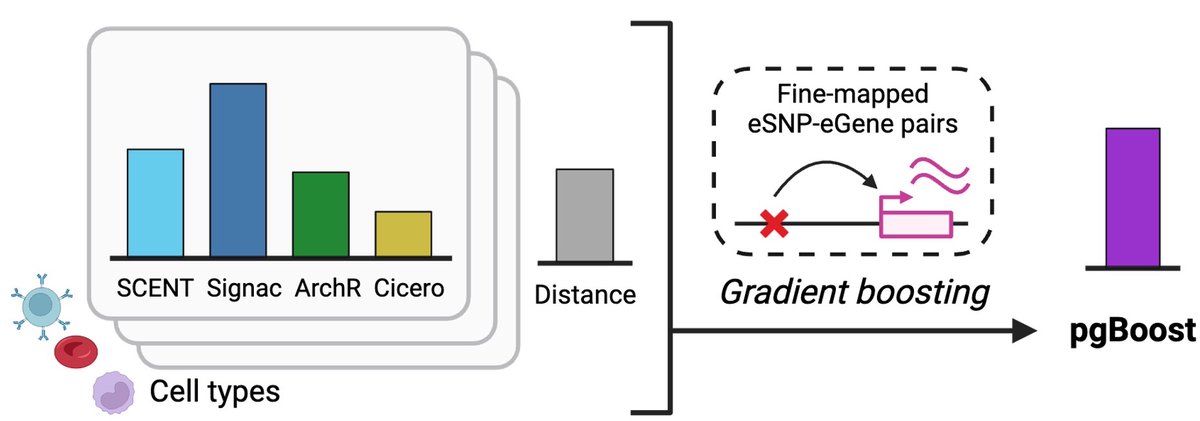
Linking regulatory variants to target genes by integrating single-cell multiome methods and genomic distance medrxiv.org/cgi/content/sh… #medRxiv

