Tweet fixado
N Molecular Systems
16 posts



(1/3) Molecular defense systems: bacteria deploy motorized spindles to entangle & degrade viral DNA.
Checkout this intriguing work!!
Structure and mechanism of Zorya anti-phage defence system.
Nature (2025)
Image for: @NMITaylorLab
Created by: N Molecular Systems | @Nth_step

English

(2/3) Molecular defense systems: bacteria deploy motorized spindles to entangle & degrade viral DNA.
Checkout this intriguing work!!
Structure and mechanism of Zorya anti-phage defence system.
Nature (2025)
Image for: @NMITaylorLab
Created by: N Molecular Systems | @Nth_step

English

(3/3) Molecular defense systems: bacteria deploy motorized spindles to trap & degrade viral DNA.
Checkout this intriguing work!!
Structure and mechanism of the Zorya anti-phage defence system.
Nature (2025)
Image for: @NMITaylorLab
Created by: N Molecular Systems | @Nth_step

English

Molecular mechanism of substrate recognition and cleavage by human γ-secretase ▼ Science
science.org/doi/10.1126/sc…
Română

"Dissecting the conformational complexity and mechanism of a bacterial heme transporter" | A passage for heme | Nature Chemical Biology, August 2023 Vol. 19 No. 8
Open Access pdf:
nature.com/articles/s4158…
EMD-14640 | 7ZDB, ...
emdataresource.org/EMD-14640
English

HEME TRANSPORTER. Decades-old enigma in bioenergetic membrane biogenesis.
Max Planck's @MPIbp @SafarianSchara et al resolved CydDC in 23+ states/conditions 2.7-3.9Å by cryoEM, probing heme structural dynamics via mutations, 20 MD simulations.
Image (2021, version B): @Nth_step

Dansk

Thank you to Max Planck @MPIbp and to Nature @nchembio for selecting the cover. It looks great in print!
Congrats to all for the great work @MPIbp @ALIFE_VU @UGent @UniKent @MiChAeL_900922 @AhmadRezaMehdi1 @franziska_f1nke @sonjawelsch @ShepherdLabKent @HummerLab @SafarianSchara
Max Planck Institute of Biophysics@MPIbp
The role of CydDC, a potential target for new antibiotics, in the bacterial energy metabolism has remained nebulous for decades. Scientists from our Institute, @ALIFE_VU, @UGent, @UniKent solved the riddle: CydDC is a heme transporter👉biophys.mpg.de/cyddc #biophysics #cryoEM
English

@SafarianSchara @MiChAeL_900922 @franziska_f1nke @AhmadRezaMehdi1 @HummerLab Truly elegant work.
"Dissecting the conformational complexity and mechanism of a bacterial heme transporter" | Nature Chemical Biology
Open Access:
nature.com/articles/s4158…
English

This one was special! Very proud of our work, despite the long journey and dealing with most hostile reviews ever received. Major credit to @MiChAeL_900922, @franziska_f1nke, @AhmadRezaMehdi1, and @HummerLab

English

Congratulations to Safarian lab at Max Planck @MPIbp !!
It was a real pleasure working with you!
@SafarianSchara @MiChAeL_900922
nature.com/nchembio/volum…
English

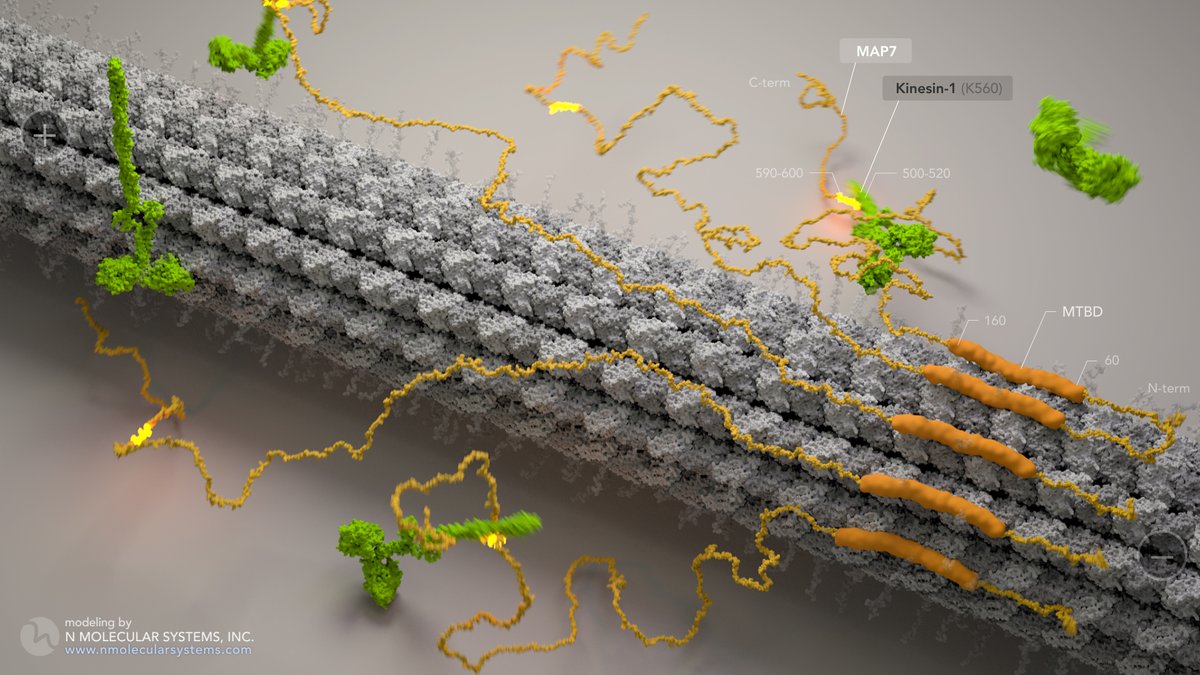
Congrats to @NogalesLab @Yildiz_Lab on insights into MAP7 & Kinesin-1 science.org/doi/10.1126/sc…
A warm flashback to our formative model (vis-a-vis 2018 "Microtubules: From Atoms to Complex Systems", EMBL) in collab with Ori-McKenney lab.
#MAPs #kinesin #microtubules #cellpolarity
English

Our collaboration on PYRD 310-helical element implicated in human disease & conserved in kinesin superfamily molecular motors, with McKenney/@ucdavisbiology @GennerichLab/@EinsteinMed Niwa/@TohokuUniPR
#AccelerateResearch, please read
advances.sciencemag.org/content/7/18/e…
@KIF1A #KAND #KIF1A
English

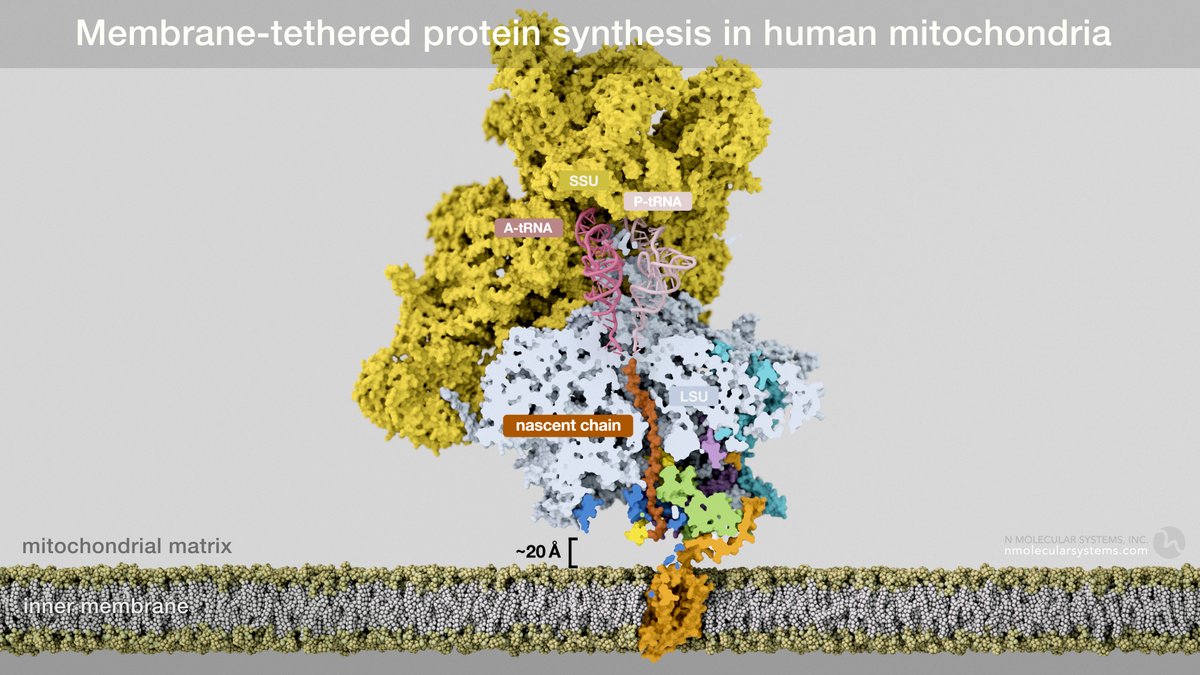
Intriguing aspect of the work @A_Amunts is 20Å gap between exit / insertase, providing for additional factors and #proteinfolding milieu (which differs from bacteria that favor early compaction of TM helices). We visualize 32 AA of #MTATP6 here (red) in place of poly-Ala in 6zm5.
English

If a machine could talk to us, what would it say?
#beautifulmachine #code #recursion #dance
youtube.com/watch?v=w0WY0p…
YouTube

English
