Chris McFarland
82 posts


Chris McFarland
@CancerEvoLab
Professional news from the Cancer Evolution Group @cwru.



Incredibly excited that our paper examining the role of ecological interactions in preexisting drug resistance is out! We tackle the apparent paradox between the cost of resistance in the absence of treatment, and the ubiquity of preexistence. 1/20 Link: go.aps.org/4c5FmH4










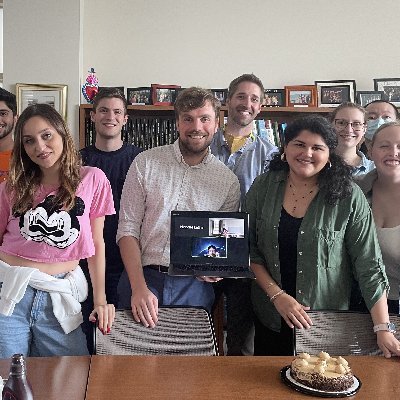
Stacey now shows cool just accepted work in @PLOSCompBiol on modeling the tumor-immune ecosystem at the tissue scale. Love the idea of using a feed forward network to make macrophage state decisions facilitating a massive speed up in computational time!


Next we have Nikolaos Dimitriou presenting, Investigating sedimentation in 3D cultures with an integrated experimental-computational framework #MathOnco23