ทวีตที่ปักหมุด
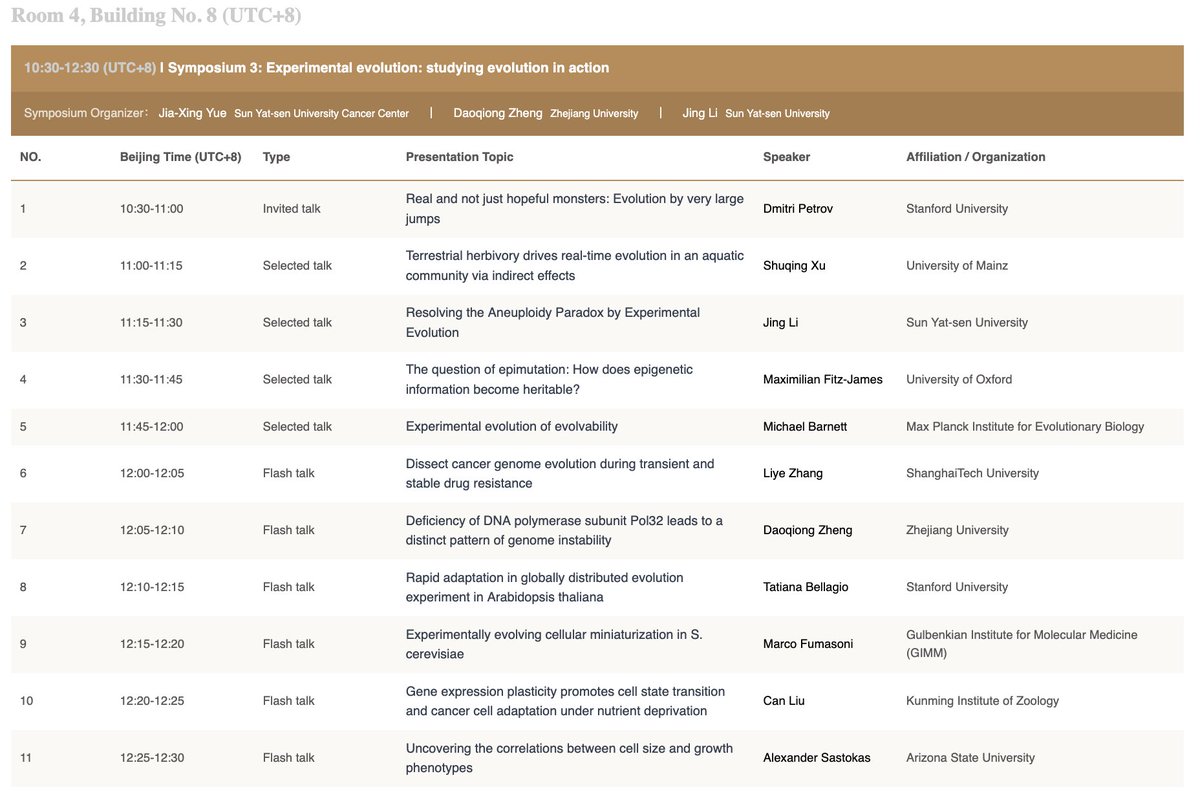
Excited to share our PNAS paper on the genome evolution of an early-diverging #amphioxus, Asymmetron lucayanum! There is a long and twisty story behind this work, which also explains the username choice of the lab PI @iAmphioxus .😀 @PNASNews
pnas.org/doi/10.1073/pn…
English