
Li Song
2.5K posts

@mourisl
毛利元光. Assistant Prof.. Bioinfo, algorithms. @BioMedDataSci. PostDoc @XShirleyLiu @lh3lh3 @dfcidatascience. PhD @JHUCompSci. https://t.co/zRfJOaf6gD #hiring



LongcallD: joint calling and phasing of small, structural and mosaic variants from long reads biorxiv.org/content/10.648… #biorxiv_genomic

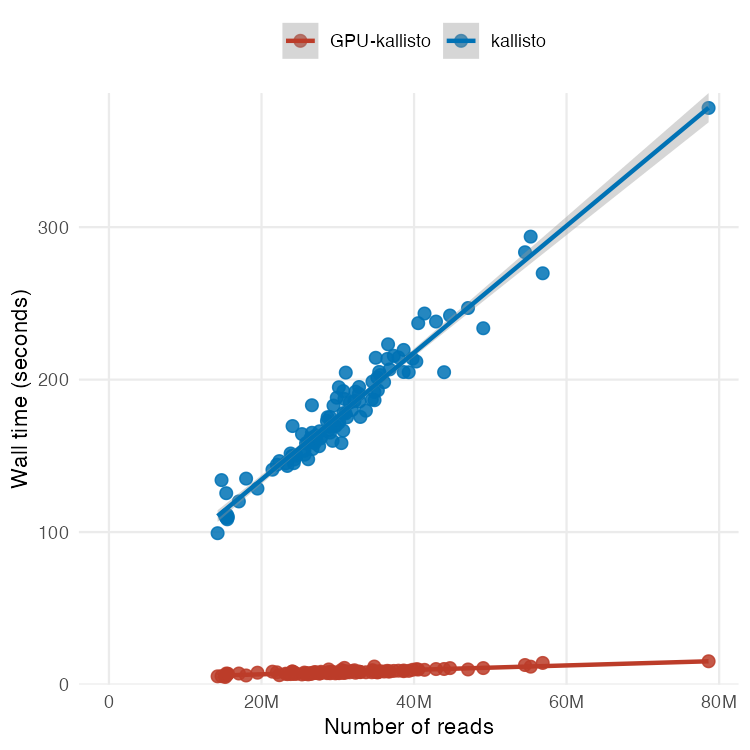



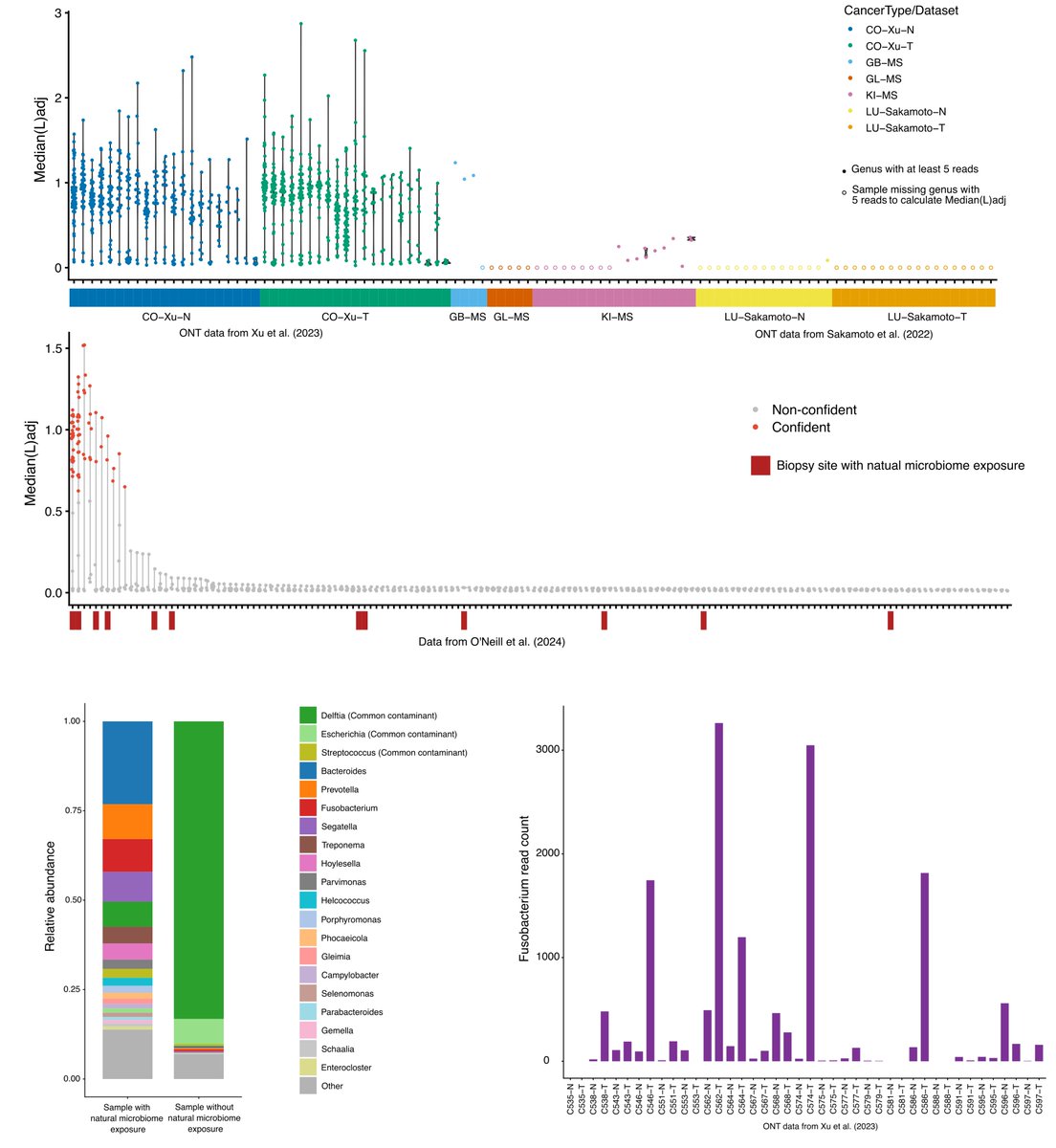


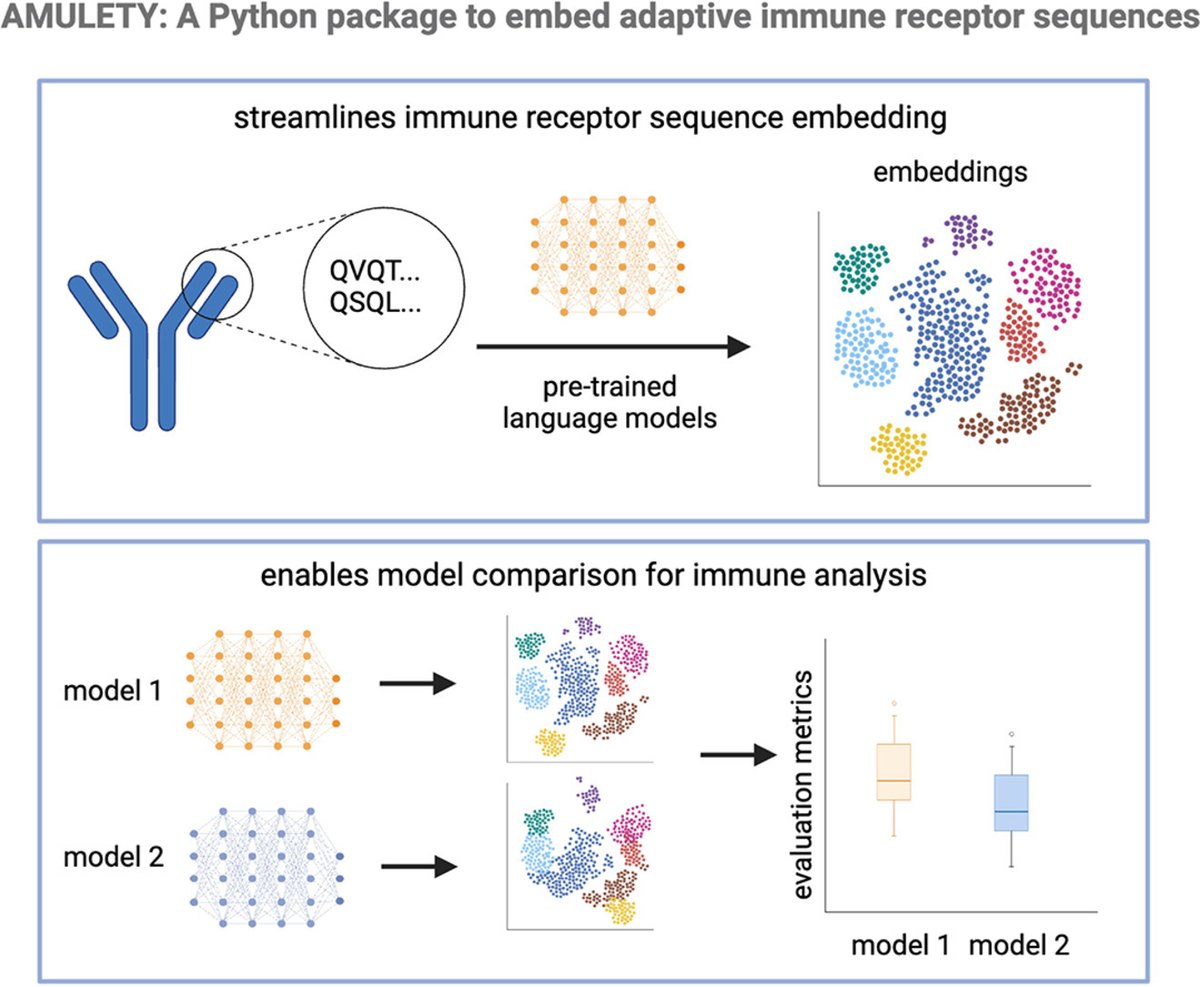


Preprint on "Improving spliced alignment by modeling splice sites with deep learning". It describes minisplice for modeling splice signals. Minimap2 and miniprot now optionally use the predicted scores to improve spliced alignment. arxiv.org/abs/2506.12986

Do you know ~60% of human SVs fall in ~1% of GRCh38? See our new preprint: arxiv.org/abs/2509.23057 and the companion blog post on how we started this project and longdust: lh3.github.io/2025/09/29/on-…. Work with @QianAlvinQin1

Literally

579 high-quality human genomes from @HumanPangenome, Arab Pangenome and individual papers (CHM13, CN1, KSA001, I002C, YAO and KOREF1). Sequences available in the AGC format (3.7GB) and FM-index in the ropebwt3 format (20.3GB). For details, see github.com/lh3/human-asm





Integrating spatial transcriptomics across platforms is hard - different gene panels, sparse data. We introduce LLOKI, using optimal transport + single-cell FMs for unified ST integration. Work led by @ellie_haber (@mldcmu) & Lane Fellow @SpencerKrieger biorxiv.org/content/10.110…


