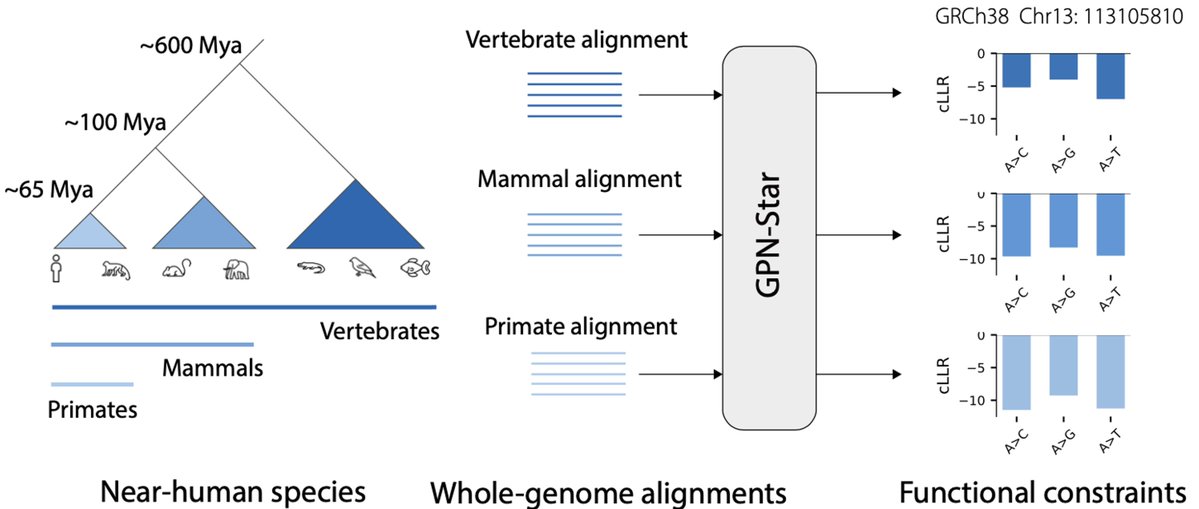
We’ve trained a multimodal AI model to turn routine pathology slides into spatial proteomics, with the potential to reduce time and cost while expanding access to cancer care.
Nicholas Ho
110 posts

@eigenNick
ML for Bio PhD Student @SCSatCMU advised by @ericxing and @jmuiuc

We’ve trained a multimodal AI model to turn routine pathology slides into spatial proteomics, with the potential to reduce time and cost while expanding access to cancer care.







Chair and Chief Executive Officer at AMD, Dr. @LisaSu, highlighted our partnership during her CES 2026 keynote. At #CES2026 in Las Vegas, GenBio AI joined @AMD to share a vision for advancing biology with AI. Together, we are building the foundation for personalized treatments made just for you. Learn more and stay tuned → x.genbio.ai/amd-genbio-ces…





📢 Submit to #ISMB2026 Proceedings! Share your latest research in #computationalbiology and #bioinformatics. Accepted papers will be published in a special open-access issue of Bioinformatics. 🗓️ Submission deadline: January 20, 2026 🔗 Learn more: iscb.org/ismb2026/call-…

Integrating spatial transcriptomics across platforms is hard - different gene panels, sparse data. We introduce LLOKI, using optimal transport + single-cell FMs for unified ST integration. Work led by @ellie_haber (@mldcmu) & Lane Fellow @SpencerKrieger biorxiv.org/content/10.110…






🚀 Xaira Therapeutics has just dropped a game-changer for AI-driven biology. Today, we unveiled X-Atlas/Orion, the largest publicly available genome-wide Perturb-seq dataset to date—spanning 8.4 million single cells with perturbations across all ~20,000 human protein-coding genes. This release is not just about scale—it’s about enabling a new era of causal, mechanistic foundation models for biology. 📝 Preprint on bioRxiv: biorxiv.org/content/10.110… 📂 Dataset on Figshare: doi.org/10.25452/figsh… 🔍 What makes X-Atlas/Orion special: 📈 Unprecedented scale & quality: Each cell profiled with deep (~16k UMIs) transcriptomics and rich metadata 🧪 Quantitative dose-response modeling: Thanks to high-fidelity sgRNA detection and ~4 guides per gene, allowing continuous modeling of genetic effects 🧬 FiCS platform: A fully industrialized single-cell perturbation system enabling rapid, reproducible experiments at massive throughput 🧠 This isn’t just “data.” It’s the biological substrate for building virtual cell models that can generalize, predict, and ultimately power AI-native drug discovery. 💬 My final take: This is a foundational moment for the field. The ability to model how genes affect cell state—quantitatively, causally, and at scale—is what we need to unlock predictive biology. Kudos to the incredible team at Xaira for open-sourcing this resource so the entire community can build on it. #PerturbSeq #SingleCell #Genomics #VirtualCell #FoundationModels #AIForBiology #Xaira #DrugDiscovery #SyntheticBiology #CausalAI More press release: Press release : 🔗 GEN article : genengnews.com/topics/artific… 🔗 BusinessWire: businesswire.com/news/home/2025…