Koichi Hashikawa
204 posts

Koichi Hashikawa
@KoichiHashikawa
Principal computational scientist, Johnson&Johnson innovative Medicine
Bay area انضم Ocak 2016
75 يتبع174 المتابعون

Super excited to introduce the RGC-ME paper nature.com/articles/s4158…! We describe the largest catalog of human protein-coding variations to date from exome data of ~1M individuals generated using a single harmonized sequencing and informatics protocol.

English
Koichi Hashikawa أُعيد تغريده

Excited to share Nicheformer! Led by @Alejandro__TL & @AnnaCSchaar, Nicheformer is a foundation model for single-cell & spatial omics. Its innovation is going beyond disassociated analysis to capture & predict local tissue context at single-cell level. biorxiv.org/content/10.110…
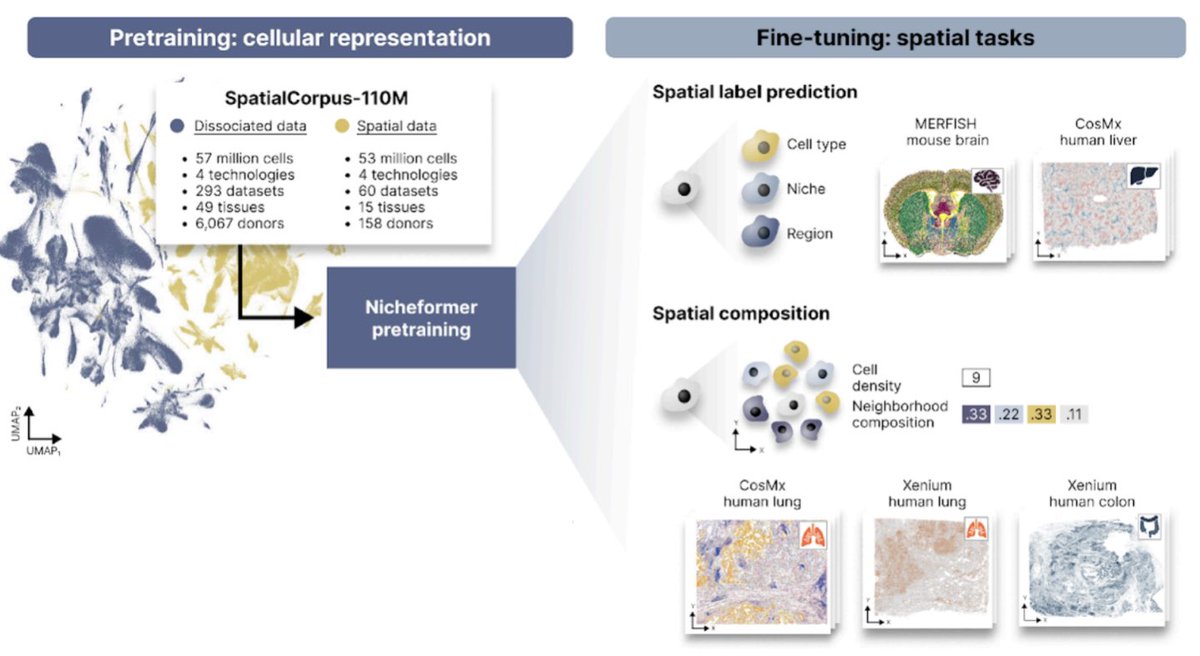
English
Koichi Hashikawa أُعيد تغريده

Very excited to announce our new preprint together with Nikolai Hecker! Dive into our findings on the conserved regulatory codes shaping cell identity across species. 🧬 [1/n]
biorxiv.org/content/10.110…
English
Koichi Hashikawa أُعيد تغريده

As my lab develops Seurat, I wanted to share some thoughts on the important Seurat and scanpy package selection preprint from @Josephmrich/@lpachter 🧵
English
Koichi Hashikawa أُعيد تغريده

Excited to share that my graduate dissertation work is now finally live on bioarxiv! Take a look if you're interested in learning more about the lateral septum's role in opioid withdrawal: biorxiv.org/content/10.110…
English
Koichi Hashikawa أُعيد تغريده

I’m so exited to share that Pixel-seq has been published! We describe a stamping method to scale production of polony gels with <1 um resolution for #spatialbiology and a clustering algorithm to segment cells based on their UMIs and transcript similarity. authors.elsevier.com/c/1g3VXL7PXiqZk
English
Koichi Hashikawa أُعيد تغريده

Here comes "spatial-ATAC-seq"!!! A wonderful collaboration between my lab at @YaleSEAS and @GoCasteloBranco at @karolinskainst
#epigenetics #spatialomics #spatialbiology
Spatial profiling of chromatin accessibility in mouse and human tissues | Nature nature.com/articles/s4158…
English
Koichi Hashikawa أُعيد تغريده

Our scCUT&Tag-pro and scChromHMM manuscript is out @NatureBiotech! Led by @Bingjie_Zhang_ and @k3yavi, we use surface proteins to integrate data from six histone modifications, RNA and ATAC - and explore chromatin state heterogeneity in single cells: nature.com/articles/s4158…
English
Koichi Hashikawa أُعيد تغريده

Our new preprint, led by @YUHANHAO2, introduces bridge integration: cross-modality alignment via a multiomic 'bridge' dataset.
We map single-cell ATAC-seq, methylation, CUT&Tag and CyTOF profiles onto scRNA-seq references- scaling to millions of cells.
biorxiv.org/content/10.110…
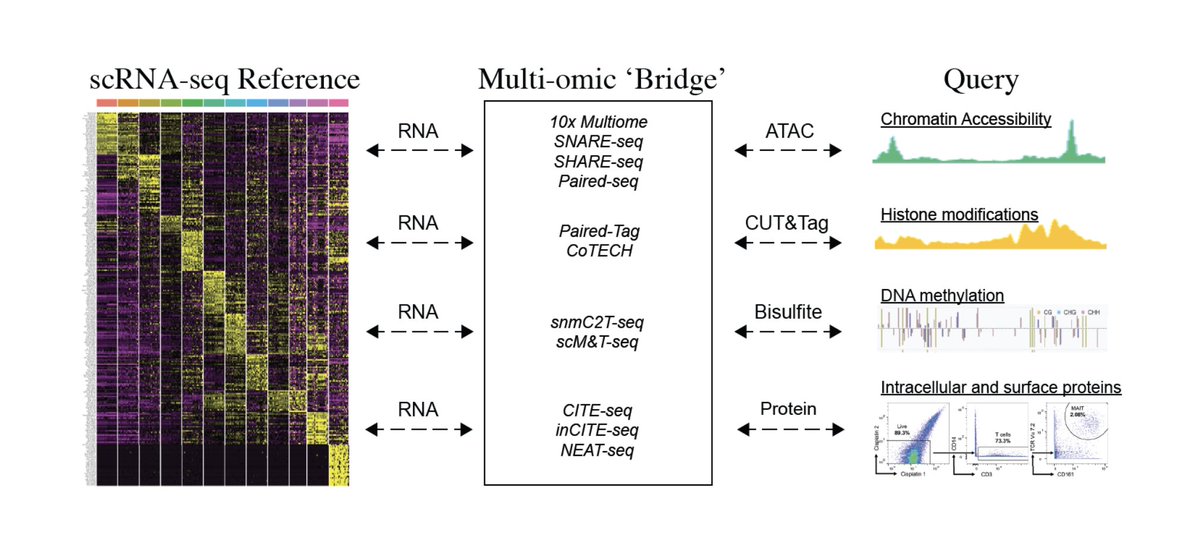
English
Koichi Hashikawa أُعيد تغريده

We are excited to introduce CaRPool-seq, a new technology that utilizes Cas13 to perform combinatorial single-cell perturbation screens! Check it out, esp. if you're interested in simultaneously perturbing multiple genes and exploring genetic interactions: biorxiv.org/content/10.110…
English
Koichi Hashikawa أُعيد تغريده

Interested in single cell genomics but need help getting started? Check out my lab's Single Cell Genomics Day on March 4. Talks will feature recent exciting computational and experimental advances and will be livestreamed at satijalab.org/scgd. Please RT/spread the word!
English
Koichi Hashikawa أُعيد تغريده

Our next @cegs_ica virtual seminar is Nov 16 at 12PM EST. @Bingjie_Zhang_ and @k3yavi will introduce scCUT&Tag-pro and scChromHMM - tools to measure, integrate, and analyze histone modification profiles in single cells.
Free registration at: nygenome.zoom.us/meeting/regist…

English
Koichi Hashikawa أُعيد تغريده

Very excited to have this out! Though this is "only" a perspective piece, I personally feel that this is my most important work yet. 1/15 authors.elsevier.com/c/1dy893BtfGzk…
English
Koichi Hashikawa أُعيد تغريده
Koichi Hashikawa أُعيد تغريده

Happy to share the last of my postdoc work. A huge thank you to the team of @gstuber @MarcusBasiri @koichihashikawa Yoshiko Hashikawa and Jill Liu (not on twitter) and collaborators at Stanford who made it possible 🧵1/6 authors.elsevier.com/c/1dtD63BtfGzk…
English
Koichi Hashikawa أُعيد تغريده

Latest and final paper from @WhoIsMarkRossi‘s time in the lab. Mark is now at @RutgersU and is recruiting. Get in touch w/him if this work interests you.
authors.elsevier.com/a/1dtD63BtfGzk…
English
Koichi Hashikawa أُعيد تغريده

Computational challenges and opportunities in spatially resolved transcriptomic data analysis
nature.com/articles/s4146…
English
Koichi Hashikawa أُعيد تغريده

Are you or someone you know interested in studying hypothalamic regulation of feeding? Do you want to do cutting edge science in an inclusive and supportive environment? Apply to work with us today!
Mark Rossi@WhoIsMarkRossi
JOB ALERT! We are hiring a postdoc to study hypothalamic feeding circuits using 2p imaging and ephys. APPLY HERE: jobs.rutgers.edu/postings/141145 more info: markrossilab.org
English
Koichi Hashikawa أُعيد تغريده

Here is link to the paper: rdcu.be/cxgbj 🐵🧠
Dan Quintana@dsquintana
Distribution of brain oxytocin & vasopressin V1a receptors in chimpanzees pubmed.ncbi.nlm.nih.gov/34482474/ "One notable difference is the lack of OXTR in reward regions such as the ventral pallidum and nucleus accumbens in chimpanzees, whereas OXTR is found in these regions in humans"
English
Koichi Hashikawa أُعيد تغريده

Want to genetically target neurons that integrate monosynaptic inputs from two upstream areas? A simple combination of AAV1 and INTRSECT makes it possible! Happy to share our new preprint. biorxiv.org/cgi/content/sh…
English