LatchBio
638 posts


LatchBio
@LatchBio
Data Infrastructure for Biology





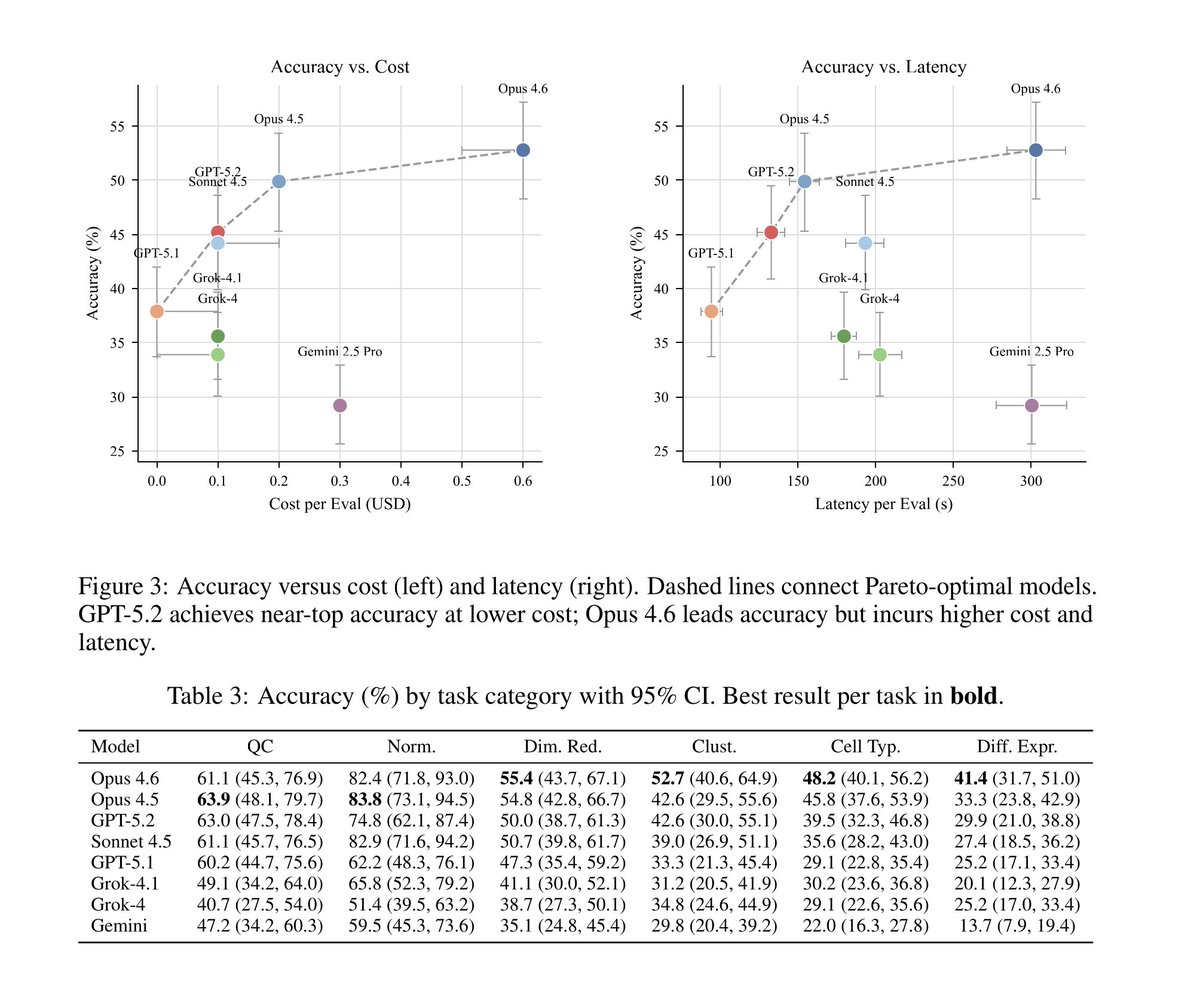
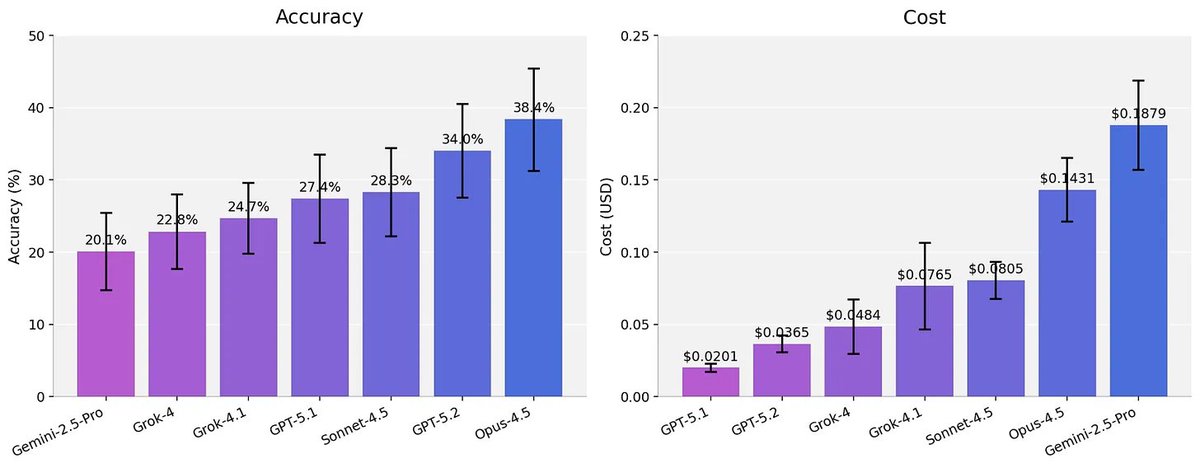
2026 will be the year of agents in biology. But we need better benchmarks. We worked with scientists to turn real world analysis into verifiable problems. SpatialBench stratifies frontier models, shows harnesses matter, and reveals distinct failure modes between model families:


2026 will be the year of agents in biology. But we need better benchmarks. We worked with scientists to turn real world analysis into verifiable problems. SpatialBench stratifies frontier models, shows harnesses matter, and reveals distinct failure modes between model families:



If you work regularly with molecular data, but don't prep it yourself, highly recommend reading the user manual for a kit you interact with cover to cover. Every little step is there for a reason + will teach you interesting biology. Small deviations at the bench often perturb your chunk of data in subtle ways.


Today, the frontier of biology research is tacit knowledge: distributed across thousands of labs in the minds of PIs/researchers. For any given disease or tissue, there are a handful of groups that know how to select, grow and identify the important molecular signatures from the models relevant to their work. A fraction of this information makes it into publication. The great task ahead of us is to formalize this knowledge into open eval banks to benchmark and improve agentic systems. If you work with spatial data and are interested in contributing to this project, please reach out.

Agents are finally starting to work in biology. We’ve partnered with Anthropic and major biotech vendors - Vizgen, AtlasXOmics, Takara, 10x Genomics - to build a tool that allows scientists to steer their own analysis with natural language. Raw spatial data to publication quality figures. Our team believes this will soon be the standard way biologists interact with data. Spatial biology agents look a bit different from coding products: - tailored to the molecular details of each kit type - run in sandboxes on very large machines - orchestrate data infra, eg. bioinformatics workflows, with tool calls - build graphical analysis notebooks to communicate results Detailed breakdown of engineering decisions, product philosophy and concrete flows follows:

Agents are finally starting to work in biology. We’ve partnered with Anthropic and major biotech vendors - Vizgen, AtlasXOmics, Takara, 10x Genomics - to build a tool that allows scientists to steer their own analysis with natural language. Raw spatial data to publication quality figures. Our team believes this will soon be the standard way biologists interact with data. Spatial biology agents look a bit different from coding products: - tailored to the molecular details of each kit type - run in sandboxes on very large machines - orchestrate data infra, eg. bioinformatics workflows, with tool calls - build graphical analysis notebooks to communicate results Detailed breakdown of engineering decisions, product philosophy and concrete flows follows:

Agents are finally starting to work in biology. We’ve partnered with Anthropic and major biotech vendors - Vizgen, AtlasXOmics, Takara, 10x Genomics - to build a tool that allows scientists to steer their own analysis with natural language. Raw spatial data to publication quality figures. Our team believes this will soon be the standard way biologists interact with data. Spatial biology agents look a bit different from coding products: - tailored to the molecular details of each kit type - run in sandboxes on very large machines - orchestrate data infra, eg. bioinformatics workflows, with tool calls - build graphical analysis notebooks to communicate results Detailed breakdown of engineering decisions, product philosophy and concrete flows follows:


Spatial FFPE ATAC-seq has arrived! AtlasXomics now brings single-cell chromatin accessibility mapping to FFPE tissue, unlocking spatial epigenomic insights from the world’s vast archives of clinical samples. Using our 5.5 × 5.5 mm chip at 10 µm resolution, we can now profile open chromatin directly in preserved tissue — maintaining spatial context. Check out our first FFPE kidney dataset on the interactive data portal, showcasing the robust performance and sensitivity of this new spatial assay. View more at: lnkd.in/e-8sTMnY #SpatialBiology #Epigenetics #CancerResearch

We’re building tools to support research in the life sciences, from early discovery through to commercialization. With Claude for Life Sciences, we’ve added connectors to scientific tools, Skills, and new partnerships to make Claude more useful for scientific work.


We’re building tools to support research in the life sciences, from early discovery through to commercialization. With Claude for Life Sciences, we’ve added connectors to scientific tools, Skills, and new partnerships to make Claude more useful for scientific work.