Ben Hitz
1.2K posts

تغريدة مثبتة

This is an infrastructure project.
Join us gopetition.com/petitions/a-un…
@anshulkundaje @ontowonka 16/16
English

@sinabooeshaghi You mean we need to go Brrrr on 1000 H100s right?
English

Precision medicine, like precision engineering, requires precise tools. To achieve Formula One-level performance with medicine, we must bring engineering principles to biology. #EngineerBiology.
English

@JD_Buenrostro Even snowflakes can be classified: its.caltech.edu/~atomic/snowcr…
English

“Every cell is a special snowflake” retweet if you agree! #teamsnowflake
nature.com/articles/d4158…
English

@arjunrajlab Still it's clear that a "type" is an ensemble of states. Types should probably be mapped to known microscopic evidence; that is mostly how they are thought of anyway. It starts to get fuzzy when you consider sorting cells on surface markers.
English

@arjunrajlab This isn't a useful definition of state because it's not precise. How many tpm difference is a different state? It is not a useful definition for type because cells have multiple functions (and function itself is slippery) and they generally are not found in all combinations
English

Since this hand-wringing continues: A cell state is a list of all molecular constituents and their parameters (one crude approximation could be the transcriptome). A cell type is the set of all cellular states that can perform a particular function (you define the function).
Arjun Raj@arjunrajlab
@arnavm1 Cell type and state are actually pretty easy to define. Time for a blog post.
English

I really don't understand what is going on. How are journals and reviewers ok with this kind of junk? The many organizations that funded this work should be outraged. And sorry for the patients whose data is trashed in this way. Scientists really ought to have some self respect!
Lior Pachter@lpachter
In this UMAP, there are arrows linking nothing to nothing (see panel d). Gornisht mit gornisht, as they say. Also drawing curves on top of a UMAP built from x-ray images by repurposing RNA velocity software that already didn't make sense is next level!
English
Ben Hitz أُعيد تغريده

@Aella_Girl @snowanddrugs2 is this because Republican-coded laws can be avoided more easier if you just have the resources to do so? Which is why dem-coded laws I consider more egalitarian.
English

@snowanddrugs2 Let's just say I'm safely breaking Republican-coded laws but I'm too scared to break democrat-coded laws
English

Fascism is rising almost nowhere in the West.
They want fewer immigrants and an end to post-2010 progressive politics. There's very little desire to invade neighboring nations for lebensraum or start goose-stepping in military parades.
Arnesa Buljušmić-Kustura@arnesa_kustura
I understand that Americans are freaking out and many are thinking about moving out of the States and I would genuinely recommend they do not do that. Fascism is on the rise literally everywhere, esp in Europe. You will not be saved by just moving away.
English

@Aella_Girl actual geneticists study the specific effect of human genetic variation on a huge variety of traits, diseases and phenotypes; see. e.g.: ebi.ac.uk/gwas/
English

@DrChrisCombs Honestly thought this was an orange cat wearing a very fancy drone hat
English

@pranamanam Aren't all these methods essentially bad? Better than random sure, but...
English

Very interesting study that shows AlphaFold3 captures a relatively global effect of mutations on PPIs by learning a smoother energy landscape, but doesn't seem to be as atomicallly fine-grained as standard MD. Could still be good for generating synthetic mutant datasets (especially when the AF3 code is open-sourced)? 🤔
Paper: biorxiv.org/content/10.110…
Results: github.com/luwei0917/Alph…

English
Ben Hitz أُعيد تغريده

One funny story about this: I spent hours creating a figure in my book explaining 0 versus 1-based indexing and closed versus right-open intervals. The illustrator thought I made a careless error in starting one from 0 and the other from 1, and changed them to match 😱
James Pitt@Sahelanth
@vsbuffalo @jgschraiber One of the many terrifying things I learned from your Bioinformatics Data Skills book!
English
Ben Hitz أُعيد تغريده

@lastpositivist the one that's a parody of heists is wonderful. Which one is that @bonscotthoughts ?
English
Ben Hitz أُعيد تغريده

Our paper "A machine readable specification for genomics assays" is now published in Bioinformatics, @OUPBioinfo. In short, we present a lightweight file format and command-line tool to document the structure of sequencing reads. Coauthored with @XiChenUoM and @lpachter.
Paper: doi.org/10.1093/bioinf…
Code: github.com/pachterlab/seq…
What is in my sequencing reads?
Sequencing machines produce text files, called FASTQs, that contain reads or sequences of DNA molecules. Assay developers and data generators deeply understand the contents of their reads; they know the location and presence biological and synthetic constructs like cellular barcodes. Collaborators, reproducers, and other scientists may not despite their sometimes obscure addition to supplementary material.
Take for example the @10xGenomics Multiome assay. The 10x Genomics documentation [1] spells out the read structure for each modality: RNA reads contain a synthetic 16bp barcode, a 12 bp "randomly" generated unique molecular barcode (UMI), as well as cDNA that was captured via polyA capture. The ATAC reads consist of genomic DNA and a 16bp cellular barcode. However the 10x Genomics website explains that the ATAC portion of the 10x Multiome data contains an little-known 8bp constant sequence spacer that proceeds the 16bp cell barcode. So saying that you have "10x Multiome" reads is a necessary but not sufficient condition to know the contents of your FASTQ reads.
The reason is because the FASTQ read structure is dependent on both the assay as well as the sequencing machine/recipe used; a sequencing library produced from one single-cell assay can yield different read structures depending these parameters. Take the the 10x Multiome assay. The ATAC 24bp barcode + spacer is usually sequenced as the i5 index read. Since the NextSeq 500/550 does not support a 24bp i5 read, the user must specify "dark cycles" (10x details the impact of this [2]) to skip the 8bp spacer. This yields a 16bp cell barcode in the i5 FASTQ file. If, however, the 10x Multiome ATAC library was not sequenced with dark cycles then the i5 FASTQ file will contain a 24bp spacer + barcode. I was originally unaware of the 8bp spacer and the use of dark cycles in Multiome library sequencing. But as I was recently looking ATAC reads I realized the impact it had on my count matrices; I had been extracting half of the cell barcode and all of the spacer. This meant I was performing barcode error correction and was UMI collapsing the, mostly similar, cell barcodes to produce few counts.
This decoupling of read structure between the sequencing machine and the assay places a high priority on documenting read structure in a sequencer and assay-specific manner so that preprocessing tools can accurately extract and process relevant sequenced elements.
A machine-readable specification
I was inspired by @XiChenUoM 's efforts (which started while in @teichlab) in documenting sequencing reads of assays and I came up with an idea to document read structure in a machine- and human-readable specification.
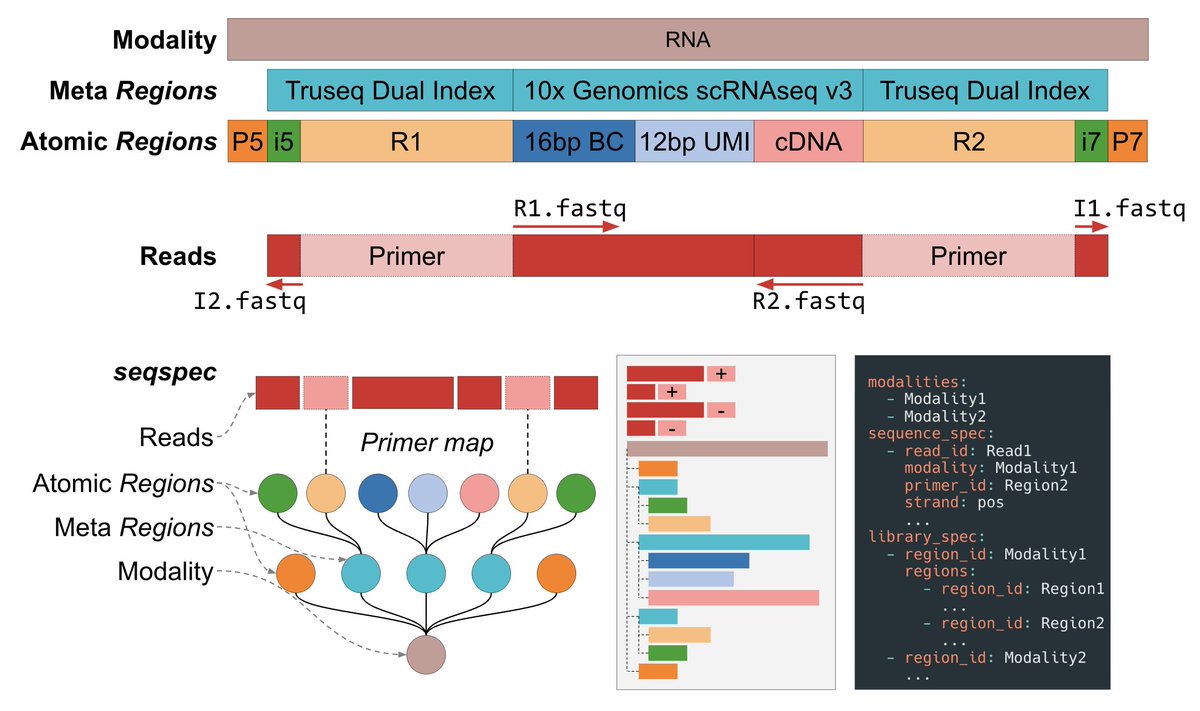
The specification is called seqspec [3]. The specification details the structure of a YAML file that allows users to specify and annotate the types of sequences that are contained in their FASTQ data. seqspec uses a nested representation of "Regions" and "Reads" that allows users to annotate groups of sequenced elements and map sequencing reads to sequencing primers. This enables, for example, all of the elements contained in Read 1 of a FASTQ file, such as the barcode and UMI in the 10x RNA assay to be annotated as belonging to Read 1. The spec also comes with an accompanying seqspec command line tool which gives users who annotate their sequencing assays many benefits:
1. Reproducibility and verifiability of the assay structure
2. Positional extraction of relevant features
3. Visualization of the sequencing structure
The seqspec command line tool makes it straightforward to extract the positional index of sequenced elements. The barcodes in the 10x Multiome dataset could have easily been identified as starting 8bp into the reads with the seqspec index command. The tool also also makes it straightforward to visualize the structure of your sequencing reads. seqspec print can produce publication-ready figure of your read structure. Most importantly, seqspec makes it easy for others to reanalyze data for which a seqspec exists, bringing about verifiability of analysis results.
seqspec adoption
seqspec aims to make genomics processing correct and reproducible. seqspec was recently adopted as the first standard in the @IGVFConsortium and we anticipate the publication of terabytes of sequencing data alongside their seqspec read annotations. I personally believe seqspec will be transformative for reproducibility and analysis efforts, in particular for those undertaken by consortia. I hope that public databases (like the @NCBI SRA/GEO and DDBJ) will test out seqspec and look to adopt it as a standard for data submission.
seqspec is freely available, open source, open to contributions, useable, and well documented. Please take a look at the GitHub repo and try it out! We welcome feedback.
[1] 10xgenomics.com/support/single…
[2] kb.10xgenomics.com/hc/en-us/artic…
[3] github.com/pachterlab/seq…
English

@RyanMarino As someone with a colonoscopy scheduled Friday I wonder the same thing
English







