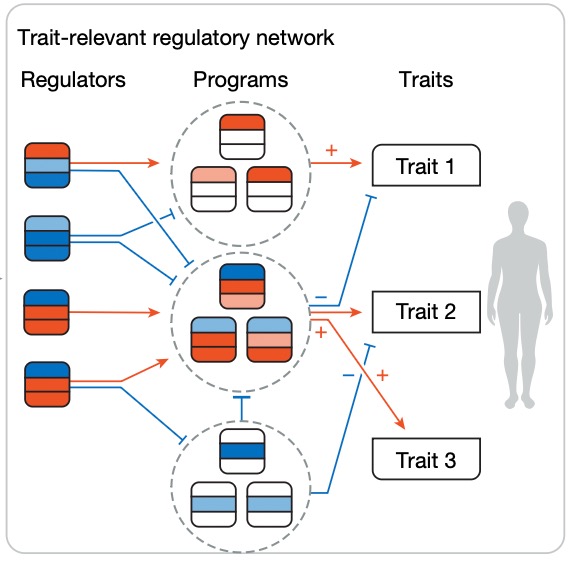
@JoeEcker @NaturePlants What a cool study! Congrats to Joseph and the whole team
English
Alexandre Marand
1.4K posts

@Marand_Lab
Assistant Professor @UMich. Chromatin biology, cis-regulatory dynamics, plant genomes.








Together with Emma Dann, we are thrilled to present a massive new Perturb-seq atlas of 22M primary CD4+ T cells, from 4 donors, across 3 timepoints – the result of a decade-long collaboration between the Marson (@MarsonLab) and Pritchard (@jkpritch) labs. 🧵👇