Alex Wolf retweetet
Alex Wolf
174 posts

Alex Wolf
@falexwolf
Open data infra for biology @laminlabs. Previously, created Scanpy & led build-up of Cellarity's compute platform.
Munich he/him Beigetreten Kasım 2016
411 Folgt2.2K Follower
Alex Wolf retweetet
Alex Wolf retweetet

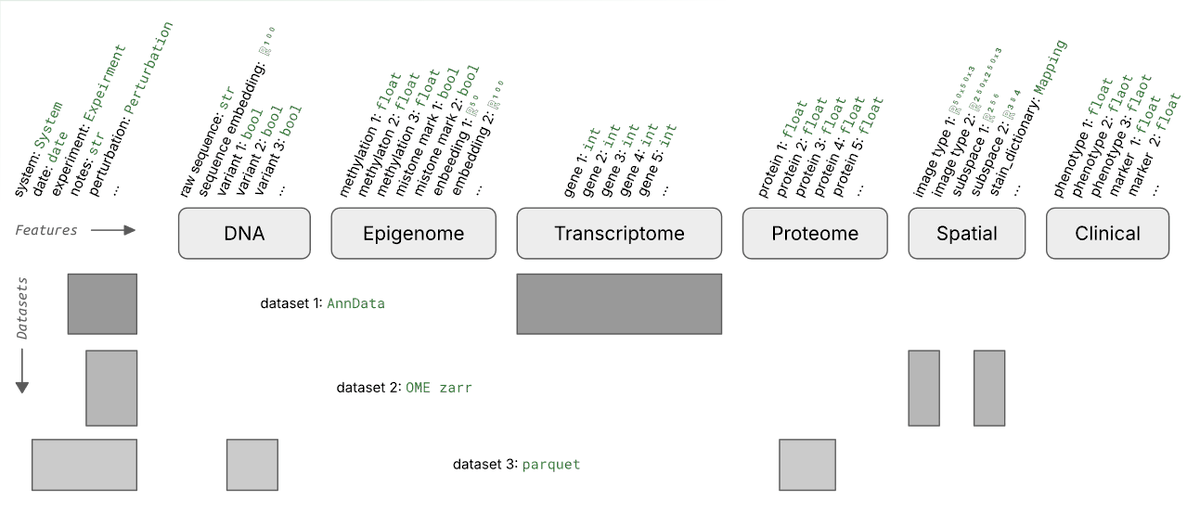
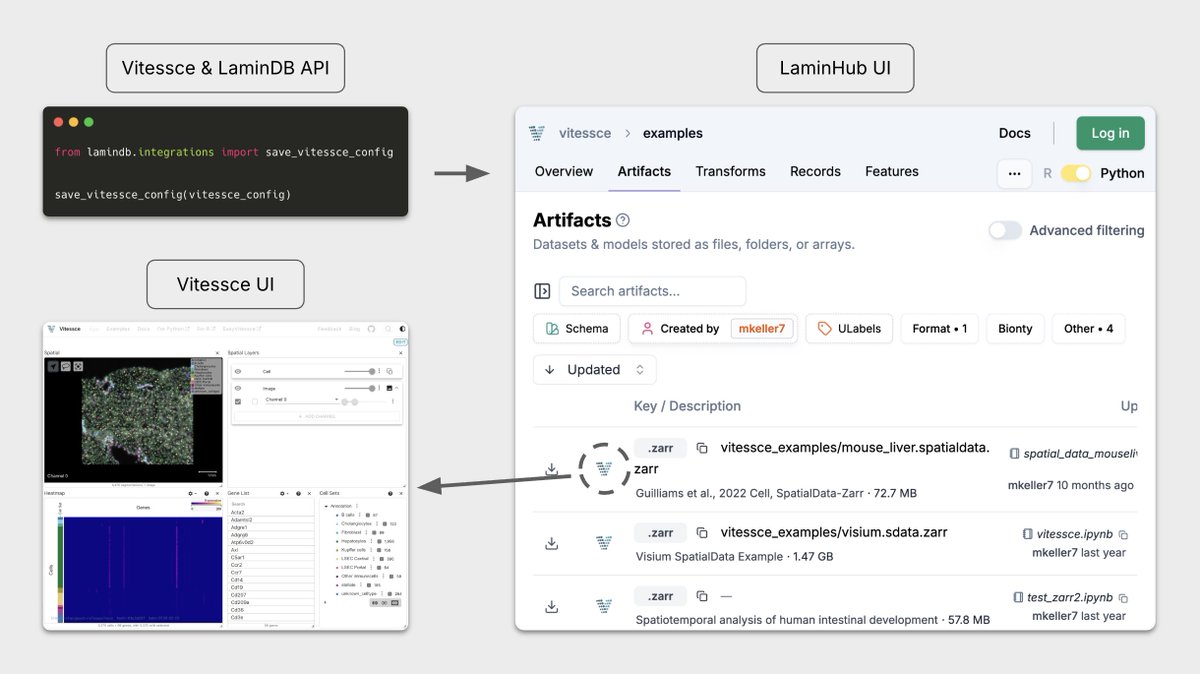
Two years ago we partnered with Mark Keller from Nils Gehlenborg’s Lab at Harvard to make Vitessce work seamlessly with LaminDB for interactive visualization of multimodal + spatial datasets.
The integration has found much use across academia, biotech, and pharma — so we wrote up on design principles & use cases.
This was a team effort involving Altana, Richard & Sunny in addition to Mark.
Read the post: blog.lamin.ai/vitessce
English

Good memory for agents.
I don't think anyone debates that this is the key bottleneck of current AI systems, so I want to spend most of this note on what an optimal "shared memory layer" for agents and humans could look like.
I think that's particularly relevant from the angle that a good part of the magic of agents is -- unlike the merely intelligent compute machine that is an LLM -- their ability to autonomously retrieve context if it's presented to them in the right way.
Agents can solve their own biggest bottleneck if "context engineering" is done right.
In this note I want to refer to the "layer" that achieves this as the "shared memory layer".
This complements the procedural side of context engineering.
The past year revealed that for now the "right way to present context to agents" seems to be files in storage paired with established API-based systems of record.
It doesn't seem to be tensors, vector databases, or RAG systems.
I found that interesting because I was never in the "symbolic camp" when it came to developing machine learning models.
So I asked Gemini 3 to clear this up: Can you disentangle why the symbolic representations in markdown notes & standard SQL databases, i.e., words and tables representing related concepts, are preferable for persisting thoughts whereas we use tensors for modeling/enabling thoughts in the first place?
English
Alex Wolf retweetet

Very good piece on the European Union and current policy problems from @lugaricano, Bengt Holmström, and @CompetitionProf: constitutionofinnovation.eu.
English
Alex Wolf retweetet

Over the past week, @arcinstitute published three new discoveries that I’m very proud of.
• The world's first functional AI-generated genomes. Using Evo 2 (the largest biology ML model ever trained, which Arc released in partnership with @nvidia in February), Arc scientists took advantage of the fact that Evo 2 is a generative model to produce completely new sequences for complete phage genomes. That is, they used AI to produce wholly new, never-before-seen-by-nature genomes. They experimentally synthesized these genomes and showed that these AI-generated phages actually work, killing E. coli bacteria with high efficacy.
• Germinal, an AI system for creating new antibodies. Antibody design is one of the great problems of medical biology given their obvious importance and usefulness for creating therapeutics. (Antibodies are tiny particles that help the immune system identify pathogens and other harmful intruders. See also the recent Works in Progress article on this topic: [1].) Today, designing effective antibodies is very expensive and slow. Germinal is a cheap and fast way to produce drug candidates, with success rates of up to 22%. This means that one can go from having to screen thousands of candidates in the lab to screening perhaps a few dozen. It's early, but I suspect that better methods for designing antibodies will be a very big deal for disease treatment in the coming years.
• Today, we published a paper showing that “bridge editing”, which Arc scientists first introduced last year, can make precise edits in human cells that are up to 1 million base pairs long, and without relying on intrinsically unpredictable cellular repair machinery (which CRISPR requires, often leading to editing mistakes). They showed that it’s possible to use this editing to cut out the DNA repeats that cause Friedreich’s ataxia (a neurological disease), an approach which should also be relevant to Huntington’s and other similar disorders. One particularly cool thing about it is that it’s possible to specify every nucleotide within the extended editing window, meaning that recursive bridge edits could potentially be a powerful way to reprogram even biological traits that are caused by many genetic mutations. (Genetic therapies today target single mutations.)
Arc is pretty new. Its doors opened in mid 2022, and it's now 300 people. I’m excited about these discoveries because they show that a number of our hopes in starting Arc are starting to pay off:
• AI/ML and computation are at the center of all three. That is obviously true for the first two, but the mobile genetic element behind bridge editing was also discovered as a result of a complex computational search. One of our premises in starting Arc was the belief that the intersection of software/AI and experimental wet lab biology should enable great things. (And besides requiring great computational work, all three of these also required strong wet lab work, tightly coordinated under a single physical roof.)
• We’ve been toying with the idea that a handful of technologies are enabling a new kind of “Turing loop” in biology: sequencing advances (including single-cell sequencing) give us new ways to read; transformers and AI gives us new ways to think; and functional genomics (such as bridge editing) give us new ways to ways to write. This trio of discoveries span each part of this loop, and we’re hopeful that there’ll be compounding returns in improving each part.
• Arc is a non-profit, which we hoped would make collaborating with others easier, since we can avoid worries about financial return. This is indeed proving important, and all three of these projects involved close partnership with others. Germinal was done in partnership with @SynBioGaoLab at Stanford; Evo 2 was trained in partnership with Nvidia. Bridge editing was jointly published with a structure from the @HNisimasu Lab at the University of Tokyo. Arc tries to make its discoveries useful (see the Evo 2 Designer[2]) for others, and the code behind the computational projects is open source, hopefully making it easy for others to spot new opportunities for collaboration and partnership in the future. Most of all, Arc itself is an ongoing collaboration with @UCSF, @UCBerkeley, and @Stanford.
• With Arc, we wanted to enable better bottom-up and top-down work. With the fully flexible, no-strings-attached funding that we provide to investigators, we want to enable completely unexpected discoveries and avenues of investigation. With our institute initiatives (around creating a virtual cell and curing Alzheimer’s), we want to bring to bear a scale and level of coordination that’s usually difficult in basic science. Germinal is a “surprise” discovery that didn’t involve top-down coordination, whereas Evo 2 is the result of ambitious high-level planning and funding.
• Humanity has never cured a complex disease (a category that includes most neurodegenerative diseases, most cancers, and most autoimmune diseases), and my hope is that Arc can help change this. It’s also clear that AI will revolutionize biology, and I hope that Arc can effectively aggregate the ingredients needed to fully capitalize on its promise. I’m biased, but I think some of the coolest biology in the world is currently being done at Arc. (They’re always hiring if you’re interested.)
While I’m a cofounder of Arc, I spend almost all my time on Stripe, where we spend our time building economic infrastructure for the internet. All credit for Arc’s progress should go to the remarkable scientists and staff who’ve made Arc their home or who’ve chosen to collaborate with us. (You can read more about these particular discoveries in these threads: [3], [4], [5].) I’m also very grateful to the amazing Stripe employees who’ve built the company that makes Arc’s ongoing work possible, and to the millions of customers who’ve chosen to partner with Stripe. John and I feel fortunate to be able to support Arc’s work to the extent that we do.
Maybe this is reading too much into it, but I sometimes feel that there’s a commonality between @arcinstitute and @stripe. Both biology and economic infrastructure involve reasoning about complex systems with many levels of emergent effects, and in both cases building the right tools can have almost unboundedly large benefits. Even though progress in both tends to take a long time, it also feels like the next five years in both will be some of the most interesting in living memory.
(If economic infrastructure is your jam, we have a whole slew of fantastic announcements coming up at Stripe Tour in New York next week. Tune in!)
English
Alex Wolf retweetet
Alex Wolf retweetet
Alex Wolf retweetet

Mario Draghi's new report on EU competitiveness doesn't mince words.
"Across different metrics, a wide gap in GDP has opened up between the EU and the US, driven mainly by a more pronounced slowdown in productivity growth in Europe. Europe’s households have paid the price in foregone living standards. On a per capita basis, real disposable income has grown almost twice as much in the US as in the EU since 2000."
"First – and most importantly – Europe must profoundly refocus its collective efforts on closing the innovation gap with the US and China, especially in advanced technologies. Europe is stuck in a static industrial structure with few new companies rising up to disrupt existing industries or develop new growth engines. In fact, there is no EU company with a market capitalisation over EUR 100 billion that has been set up from scratch in the last fifty years, while all six US companies with a valuation above EUR 1 trillion have been created in this period. This lack of dynamism is self-fulfilling."
"There are not enough academic institutions achieving top levels of excellence and the pipeline from innovation into commercialisation is weak. [...] However, while the EU boasts a strong university system on average, not enough universities and research institutions are at the top. Using volume of publications in top academic science journals as an indicative metric, the EU has only three research institutions ranked among the top 50 globally, whereas the US has 21 and China 15."
"Regulatory barriers to scaling up are particularly onerous in the tech sector, especially for young companies. Regulatory barriers constrain growth in several ways. First, complex and costly procedures across fragmented national systems discourage inventors from filing Intellectual Property Rights (IPRs), hindering young companies from leveraging the Single Market. Second, the EU’s regulatory stance towards tech companies hampers innovation: the EU now has around 100 tech-focused laws and over 270 regulators active in digital networks across all Member States. Many EU laws take a precautionary approach, dictating specific business practices ex ante to avert potential risks ex post. For example, the AI Act imposes additional regulatory requirements on general purpose AI models that exceed a pre-defined threshold of computational power – a threshold which some state-of-the-art models already exceed. Third, digital companies are deterred from doing business across the EU via subsidiaries, as they face heterogeneous requirements, a proliferation of regulatory agencies and “gold plating” of EU legislation by national authorities. Fourth, limitations on data storing and processing create high compliance costs and hinder the creation of large, integrated data sets for training AI models. This fragmentation puts EU companies at a disadvantage relative to the US, which relies on the private sector to build vast data sets, and China, which can leverage its central institutions for data aggregation. This problem is compounded by EU competition enforcement possibly inhibiting intra-industry cooperation. Finally, multiple different national rules in public procurement generate high ongoing costs for cloud providers. The net effect of this burden of regulation is that only larger companies – which are often non-EU based – have the financial capacity and incentive to bear the costs of complying. Young innovative tech companies may choose not to operate in the EU at all."
More: commission.europa.eu/document/downl….
Ursula von der Leyen@vonderleyen
Dear Mario Draghi, a year ago, I asked you to prepare a report on the future of Europe’s competitiveness. No one was better placed than you to take up this challenge. Now, we are eager to listen to your views ↓ twitter.com/i/broadcasts/1…
English

@davidsebfischer @MedUni_Wien @BockLab Congrats, David! Awesome! And that’s not even so far from here
English

I’m thrilled to announce that I will join the Institute of Artificial Intelligence at the Medical University of Vienna (@MedUni_Wien), led by @BockLab, as a tenure-track assistant professor starting December 2024! Check out our website and open positions ai4biomedicine.org
English
Alex Wolf retweetet

Categorical vs categories helps disambiguate “classes vs labels” in the sense you define it, does it?
I fear that in the original definition of “ML classification”, “classes” were indeed interchangeable with “labels”.
What can be even more confusing is that ontological hierarchies (super classes and sub classes) have nothing to do with categoricals.
The former stem from standalone ontological definitions, the latter need a definition of a measurement. For instance, you can measure values for two categoricals obtained by different annotations “organism_by_expert1” and “organism_by_expert2” and both of these draw values (“cat”, “dog) from the same ontological concept/class “animal”.
English
Alex Wolf retweetet

Thrilled to reveal @falexwolf, who originally created Scanpy, as an invited speaker at the inaugural #scverse2024 conference! Get ready to dive into his expertise on single-cell innovations.
👉 Secure your spot now at scverse.org/conference2024…!

English
Alex Wolf retweetet

Exciting news for the Python Single-Cell Community!
Save the date for the first scverse conference in Munich from September 10-12. Inspiring talks, interactive workshops, and networking opportunities. Stay tuned for the registrations, speaker announcements, and more! #scverse2024

English

The pragmatic question is whether there could be CI running on both Seurat & Scanpy that monitors metrics across versions the community agrees on: @scverse_team @ivirshup @fabian_theis @satijalab
@Josephmrich
Along with documentation that explains caveats and differences.
Did the extensive work on scRNA-seq best practices in more recent years ever get fed back into Scanpy defaults (e.g., by @MDLuecken @LukasHeumos)?
If not, Scanpy's defaults are likely still in sync with the Seurat version that inspired them in 2017. 😅
github.com/scverse/scanpy…
English

As my lab develops Seurat, I wanted to share some thoughts on the important Seurat and scanpy package selection preprint from @Josephmrich/@lpachter 🧵
English

Great, Rahul! You use much better words to convey the nature of clustering.
> a major challenge in scRNA-seq analysis is that cells cannot always be perfectly grouped into discrete bins
> not deterministic, and different random seeds will affect downstream analysis. But as @_canergen has pointed out, true biological signal persists across runs with different random seeds.
x.com/falexwolf/stat…
English