Andrew A. Borkowski retweetet
Andrew A. Borkowski
489 posts



Out of everything I saw at CES this is the gadget I want the most.
How about you?
NVIDIA Data Center@NVIDIADC
Here's a closer at the new NVIDIA Project DIGITS personal AI supercomputer. Live from the #CES2025 show floor -- attendees got the first look. Learn more now. ➡️ nvda.ws/3WgYvQT #NVIDIAGraceBlackwell
English

Based on numerous requests, we are providing the open ShareIT link for UNI and CONCH. Please access it below:
Open ShareIT Read Links:
UNI: rdcu.be/dBMgh
CONCH: rdcu.be/dBMf6
Journal Links for complete pdf:
UNI: nature.com/articles/s4159…
CONCH: nature.com/articles/s4159…

English

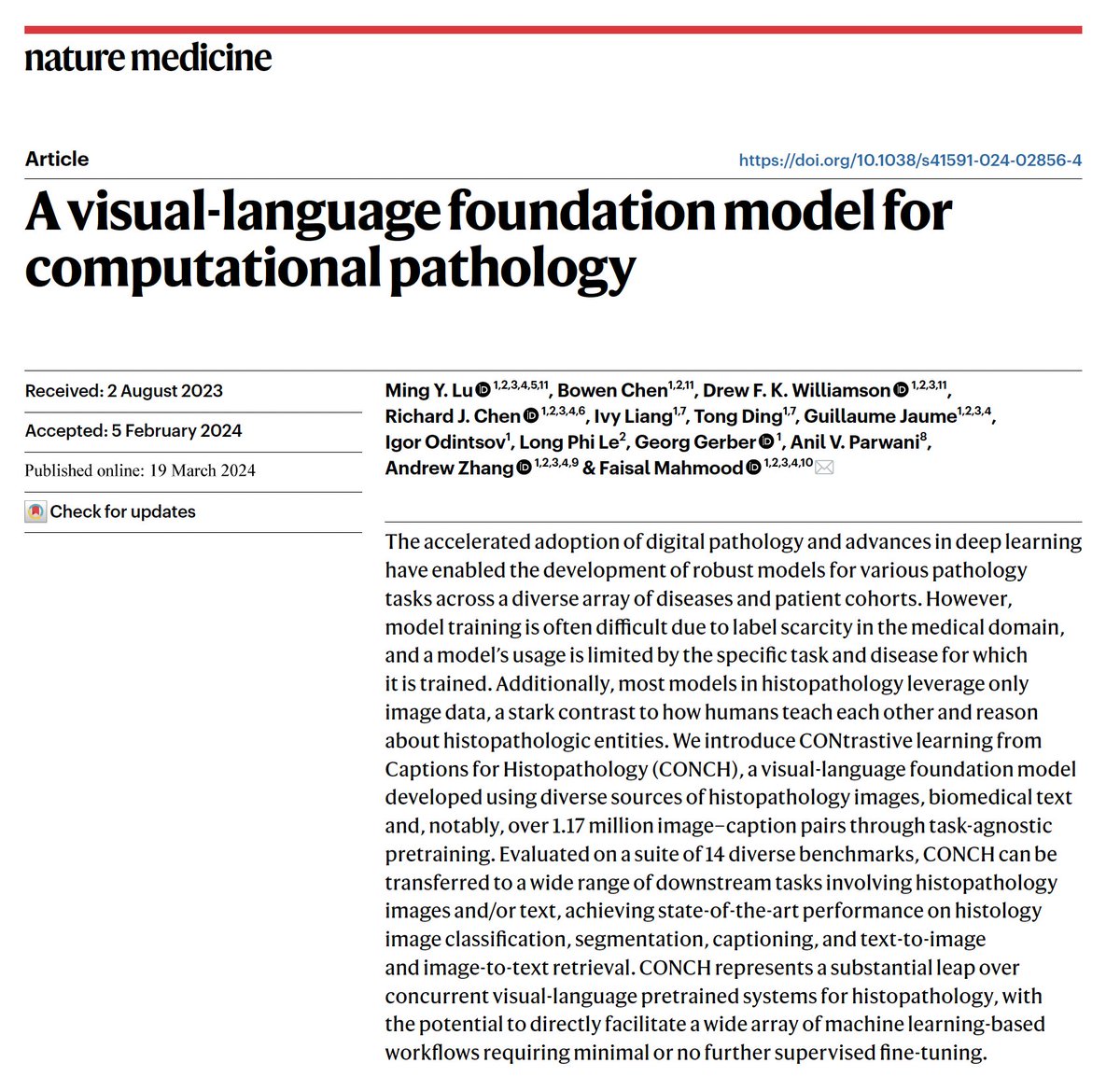
⚡️🔬📣Excited to share our two new @NatureMedicine articles, we develop computational pathology foundation models,
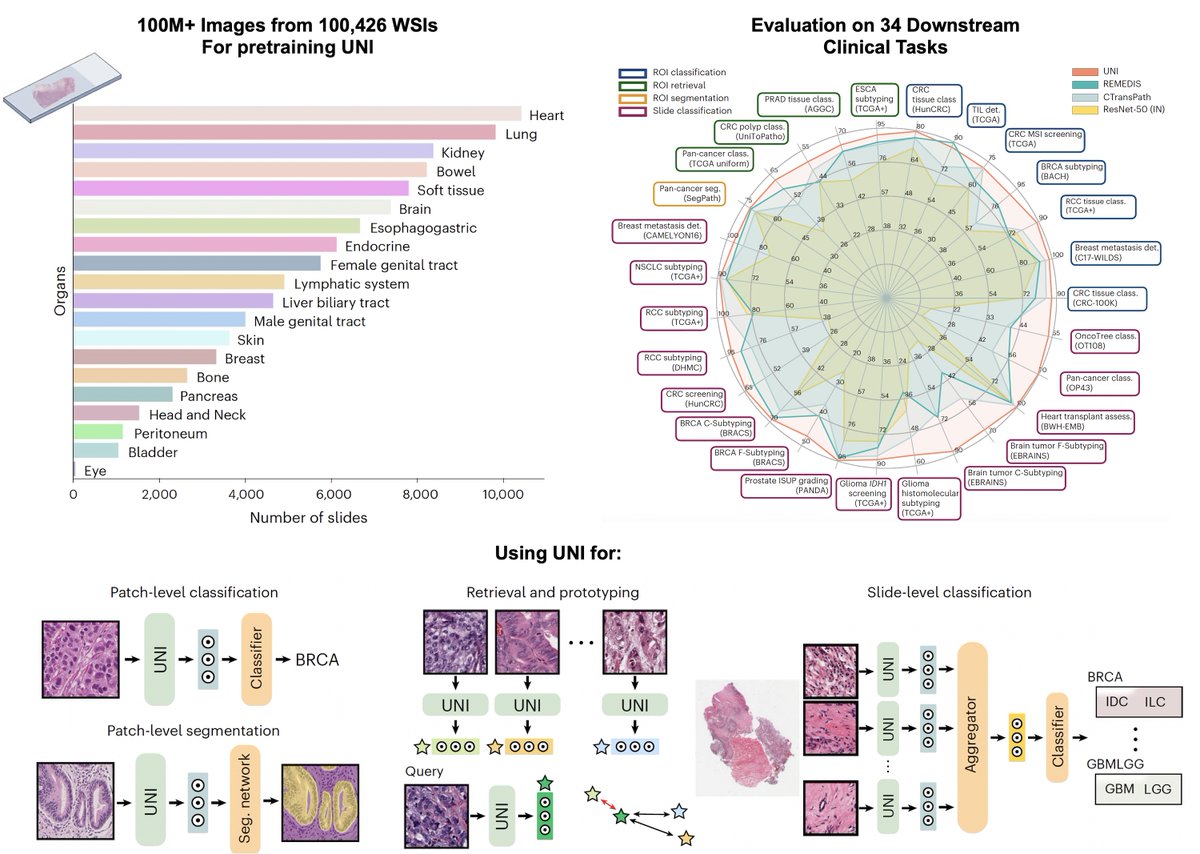
1. UNI, a self-supervised computational pathology model trained on 100 million pathology images from 100k+ slides.
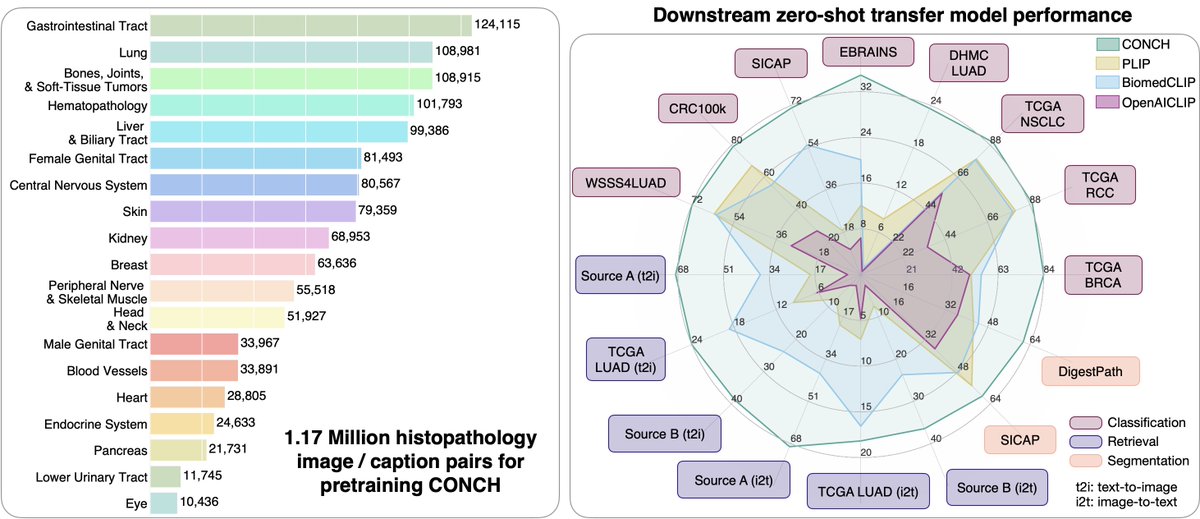
2. CONCH, a vision-language model for computational pathology trained on 1.17 million pathology image-text pairs.
Access the articles @NatureMedicine
UNI: nature.com/articles/s4159…
CONCH: nature.com/articles/s4159…
Access the code, models:
UNI: github.com/mahmoodlab/UNI
CONCH: github.com/mahmoodlab/CON…
Interesting aspects:
- Both models are evaluated on a host of different clinically relevant tasks for WSI classification, ROI classification, segmentation, image retrieval, image-to-text retrieval, text-to-image retrieval, in 0-shot, few-shot and supervised settings. These adaptations encompass the utility of large public datasets and evaluations on independent test cohorts.
- Both models exclude commonly used public computational pathology benchmarks from pre-training allowing for a much more holistic evaluation.
Some limitations: Both UNI and CONCH represent early developments in foundation models for pathology. More data, and additional evaluation is needed to realize the full potential of these models. Nevertheless, we show the models capabilities on a variety of different benchmarks with several demonstrating state-of-the-art performance.
Future work and insights: While these developments are exciting, they represent work we did about a year ago when the pre-prints were made available, since then we have been busy collecting significantly larger datasets and hope to make larger models available in the future. We have also used UNI and CONCH as the backbone for our Pathology specific chatbot, PathChat (arxiv.org/abs/2312.07814), which is further trained on hundreds of thousands of pathology specific Q-A instructions.
We are also excited to see foundation models for several other areas of biomedicine including for single cell data (nature.com/articles/s4159…), radiology (nature.com/articles/s4225…) and the general trajectory towards general purpose AI for biomedicine.
Congratulations to our superstar leaders @richardjchen @MYLu97 @DFKW_MD @TongDing99, Bowen Chen and everyone else who contributed to these studies @GuillaumeJaume @GreatAndrew90 @sharifa_sahai @Aparwani_dpath and others.


English

@AI4Pathology @NatureMedicine Fantastic work, congratulations Faisal!
English

@USFpathology Congrats and welcome to our Pathology family!
English

Congratulations to our new residents! We are proud to have you join the USF family! #USF #Pathology #Match2024 🐮🤘🏻

English

I passed my PhD Defense!!!
I am forever grateful to @Summerin3D @VirtuallyJon @ielnaqa, my amazing committee members, and Dr. Andrew Borkowski for chairing! Also thank you to Dr. Doug Ivancsits and the entire @usfradiology team for supporting me in this crazy venture ❤️

English

Happy to share our latest paper by @ElNahhasOSM et al., which was just published in @NatureComms! This work presents a new method which improves the prediction of biomarkers from routine pathology slides. nature.com/articles/s4146… @tudresden_de @katherlab @Medizin_TUD

English

@doctor_mayfield @arxiv @PennyLaneAI @ielnaqa @ml4onco @USFResearch @USFHealth @EngineeringUSF @usfradiology Great work, congrats John!
English

⚛️I have studied #QuantumComputing for a few years now and just released my #QML project on @arxiv in preprint:
arxiv.org/abs/2401.12132
3 QCNNs/VQCs + LSTM layer for binary classification of MS disability vs. VGG-RNN and ViViT.
Big news to come!
@PennyLaneAI
#phdlife #quantum

English
Andrew A. Borkowski retweetet

Hope you’ll enjoy reading our new article on #digital #pathology #Genarative #AI @MoffittNews @MoffittResearch @ml4onco @TheUSCAP @LIjournal @SIIM_Tweets #RSNA23 #pathtwitter @USFResearch @CAPA_comm @dpatweet

English
Andrew A. Borkowski retweetet

Excited to announce Med-Flamingo, a new multimodal few-shot learner specialized for the medical domain!
Last week, we uploaded the pre-print, now it’s finally live!
Paper: arxiv.org/abs/2307.15189
Code: github.com/snap-stanford/…
Model: huggingface.co/med-flamingo/m…
A short 🧵.
1/n


English

Excited to announce the publication of “Opportunistic detection of type 2 diabetes using deep learning…” co-authored with @AyisPyrros, @mattlungrenMD, @judywawira et al. in @nature journals! 🧵 #deeplearning #radiology #healthcare

English

@doctor_mayfield @RSNA @Summerin3D @VirtuallyJon @usfradiology @EngineeringUSF @ml4onco @MoffittNews @TampaVA Congratulations John!
English

The official one came in today! Very honored to receive the @RSNA #Roentgen #radres Research award as an R3.
Thank you again @Summerin3D
@VirtuallyJon @usfradiology @EngineeringUSF @ml4onco @MoffittNews @TampaVA !!!
Wait until you see what is next 🧠
#phdlife
#physicianeer


English

@doctor_mayfield @RSNA @Summerin3D @VirtuallyJon @ielnaqa @usfradiology @USFHealth @ml4onco @MoffittNews Congratulations John, well deserved!!!
English

Incredibly humbled to receive the @RSNA #Roentgen Research Award as an R3. Still stunned. Thank you to my mentors and my family who helped to support me in my many crazy endeavors @Summerin3D @VirtuallyJon @ielnaqa @usfradiology @USFHealth @ml4onco @MoffittNews
#radres #phdlife

English
Andrew A. Borkowski retweetet

Lastly (for now), Drs. Seifert, Gorlin, and @tampapath from @UFMedicine and @USFHealthMed provide a review of #AI and clinical #cytometry. This is the clearest explanatory overview of the topic I've ever seen, great for #hemepath folks new to topic.
labmed.theclinics.com/article/S0272-…
English
Andrew A. Borkowski retweetet

@isafulf @hwchase17 @realSharonZhou 2/Building Systems with the ChatGPT API: Go beyond individual prompts, and learn to build complex applications that use multiple API calls to an LLM. Also learn to evaluate an LLM's outputs for safety and accuracy, and drive iterative improvements. learn.deeplearning.ai/chatgpt-buildi…
English