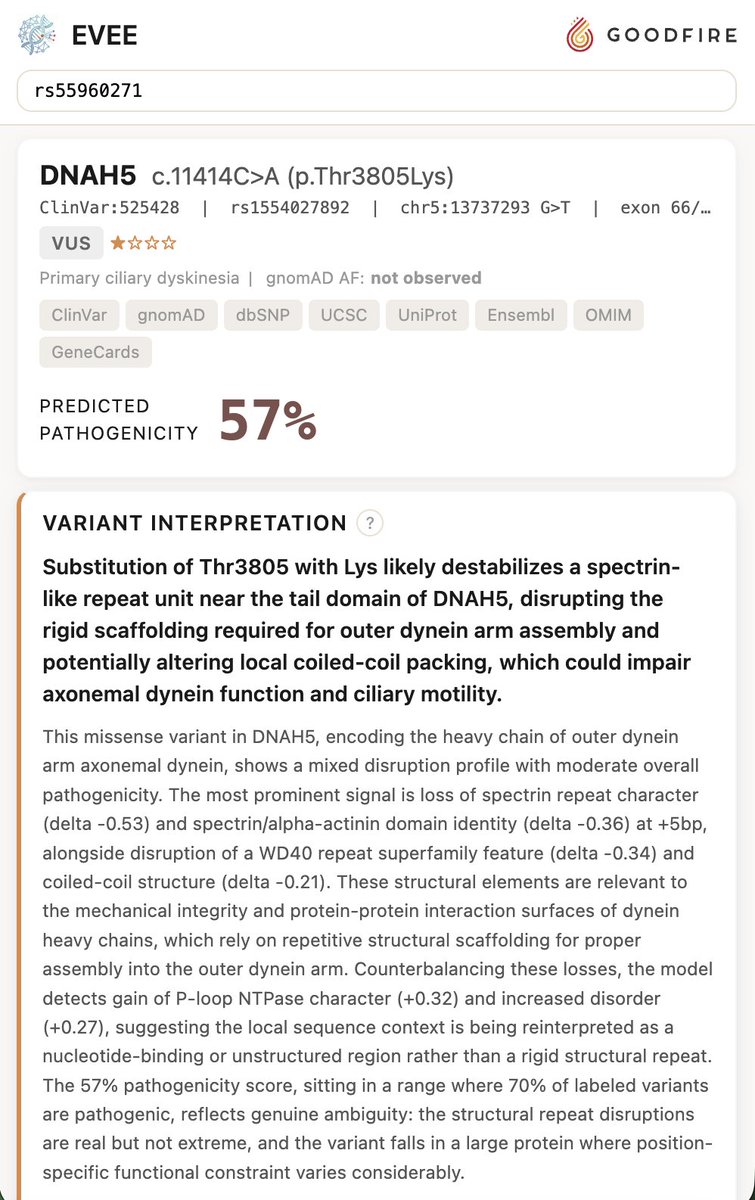
We achieved state-of-the-art performance in predicting which of 4.2 million genetic variants cause diseases by interpreting a genomics model, in a new preprint with @MayoClinic. We're now releasing an open source database for all variants in the NIH's clinvar database. 🧵(1/8)