Ryan Chow retuiteado

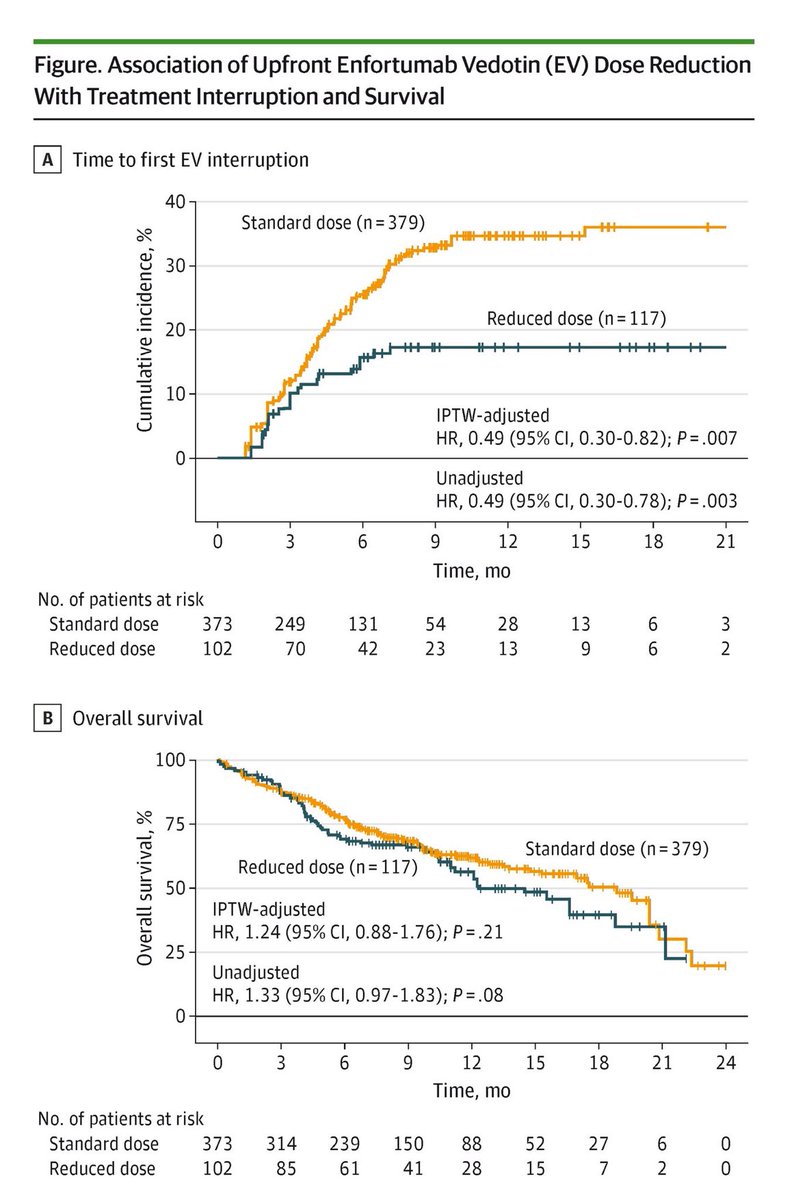
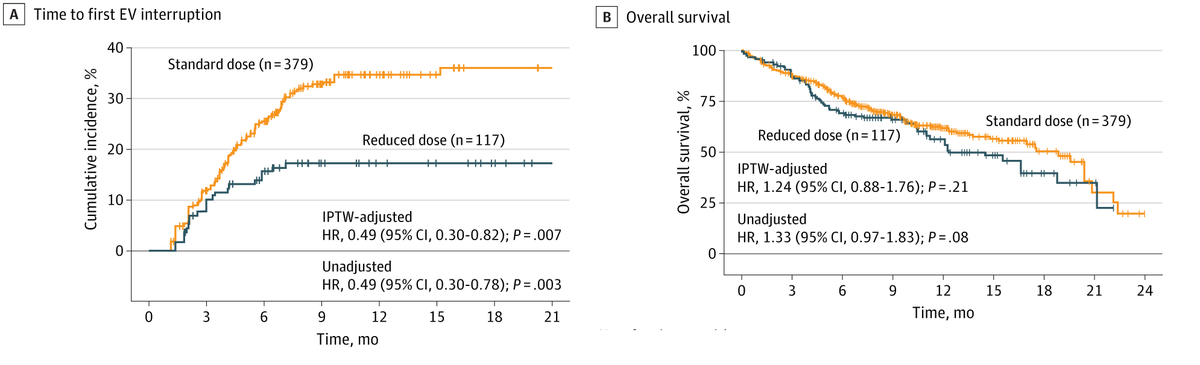
Real-World Analysis Supports Rethinking Upfront Dose Intensity of Enfortumab Vedotin in Advanced Urothelial Cancer @ScienceChow @PennCancer @Ron_cology @ramsedhom #blcsm hubs.ly/Q041HfMF0
English
Ryan Chow
106 posts

@ScienceChow
Heme/onc fellow @PennCancer | @YaleMed MD/PhD w/ @sidichen | Harvard '16 | https://t.co/ynj5I1RhPY





















#DeepLearning models have been developed to predict missense variant pathogenicity -- but how well do these models perform in a real-world clinical setting? Thrilled to share our latest in npj Precision Oncology! @Nature_NPJ | nature.com/articles/s4169… (1/8)




