Stanic Lab
1.9K posts

Stanic Lab
@StanicLabUW
Physician scientist - Immunology of Reproduction. Clinical REI. Tech and algos push sci boundary. Re-tweet not endorsement. Comments are personal opinions.





Judging by my tl there is a growing gap in understanding of AI capability. The first issue I think is around recency and tier of use. I think a lot of people tried the free tier of ChatGPT somewhere last year and allowed it to inform their views on AI a little too much. This is a group of reactions laughing at various quirks of the models, hallucinations, etc. Yes I also saw the viral videos of OpenAI's Advanced Voice mode fumbling simple queries like "should I drive or walk to the carwash". The thing is that these free and old/deprecated models don't reflect the capability in the latest round of state of the art agentic models of this year, especially OpenAI Codex and Claude Code. But that brings me to the second issue. Even if people paid $200/month to use the state of the art models, a lot of the capabilities are relatively "peaky" in highly technical areas. Typical queries around search, writing, advice, etc. are *not* the domain that has made the most noticeable and dramatic strides in capability. Partly, this is due to the technical details of reinforcement learning and its use of verifiable rewards. But partly, it's also because these use cases are not sufficiently prioritized by the companies in their hillclimbing because they don't lead to as much $$$ value. The goldmines are elsewhere, and the focus comes along. So that brings me to the second group of people, who *both* 1) pay for and use the state of the art frontier agentic models (OpenAI Codex / Claude Code) and 2) do so professionally in technical domains like programming, math and research. This group of people is subject to the highest amount of "AI Psychosis" because the recent improvements in these domains as of this year have been nothing short of staggering. When you hand a computer terminal to one of these models, you can now watch them melt programming problems that you'd normally expect to take days/weeks of work. It's this second group of people that assigns a much greater gravity to the capabilities, their slope, and various cyber-related repercussions. TLDR the people in these two groups are speaking past each other. It really is simultaneously the case that OpenAI's free and I think slightly orphaned (?) "Advanced Voice Mode" will fumble the dumbest questions in your Instagram's reels and *at the same time*, OpenAI's highest-tier and paid Codex model will go off for 1 hour to coherently restructure an entire code base, or find and exploit vulnerabilities in computer systems. This part really works and has made dramatic strides because 2 properties: 1) these domains offer explicit reward functions that are verifiable meaning they are easily amenable to reinforcement learning training (e.g. unit tests passed yes or no, in contrast to writing, which is much harder to explicitly judge), but also 2) they are a lot more valuable in b2b settings, meaning that the biggest fraction of the team is focused on improving them. So here we are.

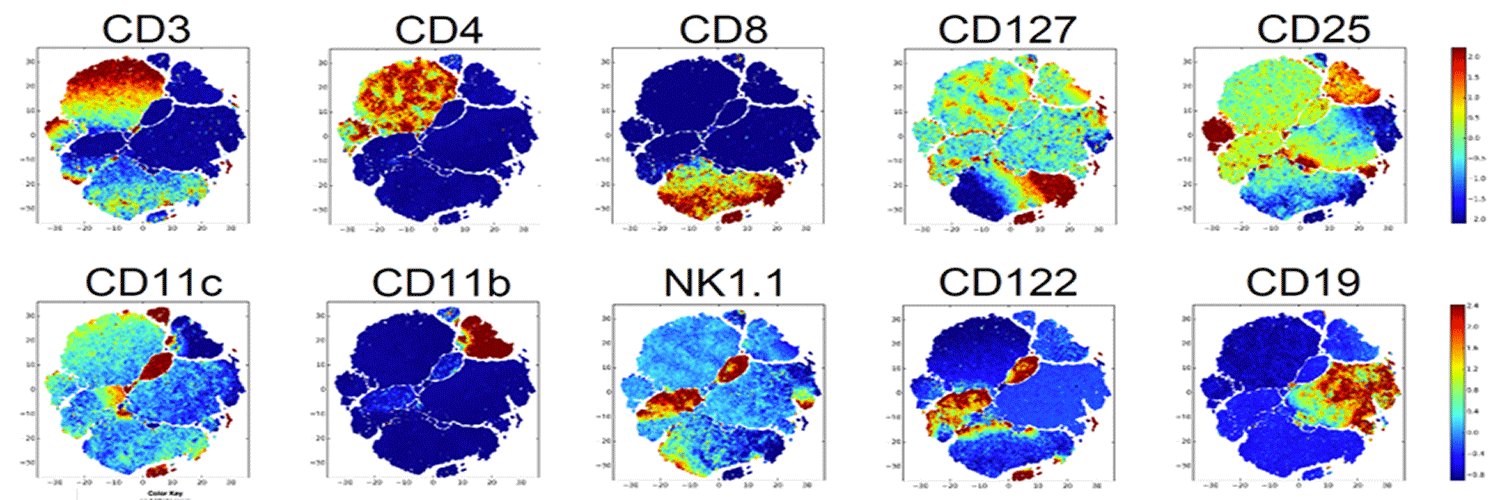
Scalable single-cell total RNA sequencing unifies coding and noncoding transcriptomics go.nature.com/4tjA7MI










@nummanali tmux grids are awesome, but i feel a need to have a proper "agent command center" IDE for teams of them, which I could maximize per monitor. E.g. I want to see/hide toggle them, see if any are idle, pop open related tools (e.g. terminal), stats (usage), etc.

We just added /btw to Claude Code! Use it to have side chain conversations while Claude is working.









