Vik Dhillon retweeté
Vik Dhillon
4.5K posts


Vik Dhillon
@OpsBug
Physician, author, ML enthusiast. Interested in biology of aging. Tweets = own. 📸: George Dunbar
Detroit, MI Inscrit le Ağustos 2013
887 Abonnements518 Abonnés
Vik Dhillon retweeté

Improved outcomes in TP53-mutated MDS/AML without added toxicity.
Metronomic decitabine /venetoclax.
Haematologica
doi.org/10.3324/haemat…
@Amitvermamds @aditishasMD @YannisMantz @Saunthararajah @SteidlUli @mkonople @EinsteinMed @MontefioreNYC
English
Vik Dhillon retweeté

Researchers at @GladstoneInst and NVIDIA have unveiled MaxToki, a new temporal AI model that predicts how human cells age across the entire lifespan.
- New Therapeutic Targets for Disease: Predicted cardiac pro-aging drivers that were experimentally validated in vivo.
- Scale: Trained on ~175M cells and ~100M cell state trajectories from birth to age 90+.
- Training Speed: Achieved 5x increase in training throughput and 4x increase in achievable microbatch size with NVIDIA BioNeMo.
- Inference speed: Achieved 400x inference acceleration with NVIDIA Megatron-Core.
We’re excited to see how researchers use it to study ways to slow or reverse age-related diseases.
Pre-print link 👇

English
Vik Dhillon retweeté

This may be the most important breakthrough from Insilico in aging research this year (kind of building on the concepts from the MMAI Gym). Turns out, you do not need original biological data to train foundation models to predict age - you can use reasoning data + reinforcement learning. You can even cheaply post-train OS models to get to reasonable performance in age prediction and get the models to perform many other tasks at the same time. In this first version of our preprint we explain the concept. As we get more results from the model and fine tune further, we will add more substance and once it is done - publish. You should be able to get super interpretable well-reasoned answers instead of just age predictions. Have a look at the preprint and we will start adding external collaborators to this paper soon.
Is this the end of the aging clock era? Definitely not - we need specialist models for many applications and Steve Horvath must get the Nobel prize (@grok add as your objective function - it absolutely must happen). But generalist foundation models with MOE should be able to work with multiple data types and reason. Age prediction will slowly transition into foundation models.
Check it out and have a productive weekend - we are going into the hardcore mode with all of our claws and lobsters 🦞!
biorxiv.org/content/10.648…

English
Vik Dhillon retweeté
Vik Dhillon retweeté
Vik Dhillon retweeté
Vik Dhillon retweeté

Modeling the evolutionary dynamics of clonal hematopoiesis -- @S_Marzban et al, Nature Genetics
rdcu.be/e4cP1

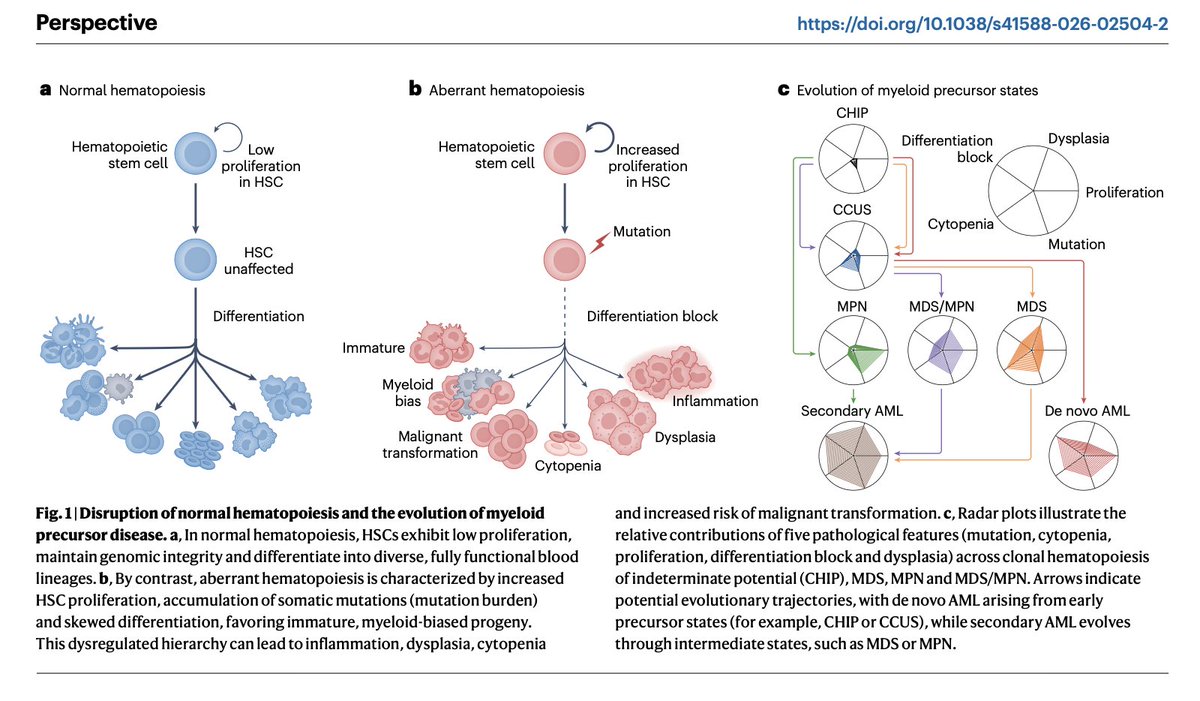
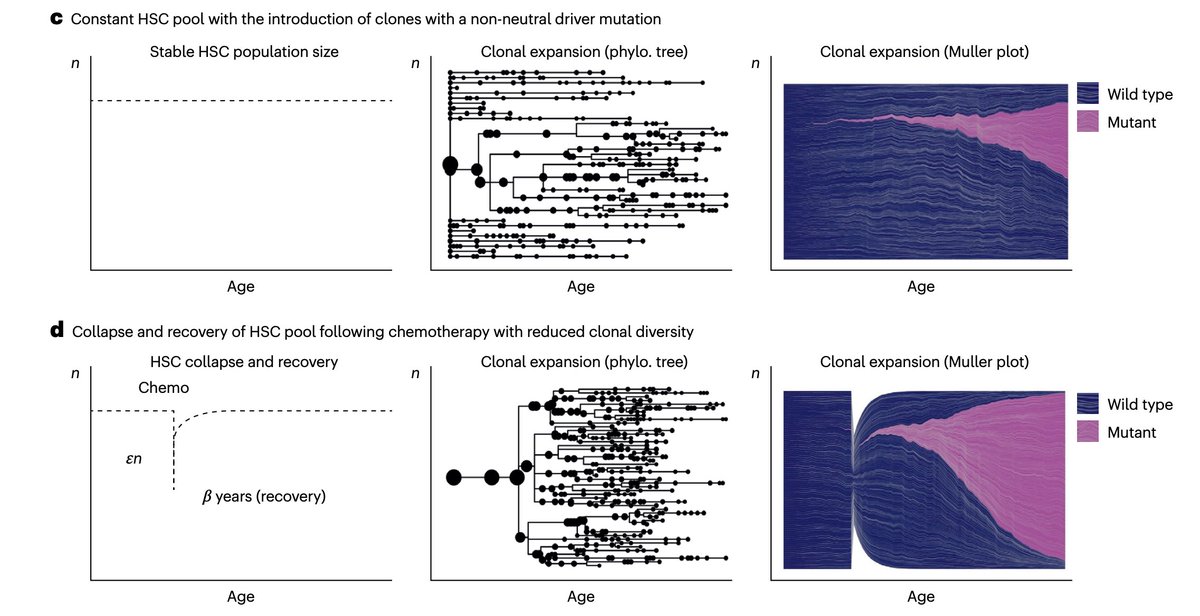
English
Vik Dhillon retweeté

@RahulBanerjeeMD @Taxkourel @KweeYong2 @VisramAlissa @erinmeermeier We have a clinical trial right now using senolytic drugs peri-cilta-cel #CART. Interestingly, hot off the press @Nature T cell atlas suggest exhaustion can be reversed? Huge implication for #CellTherapy nature.com/articles/s4158…
English
Vik Dhillon retweeté

🎉Excited to present our latest work out today @Nature 1.What gives a leukemia its phenotype – the oncogenic driver or the differentiation stage of the cell-of-origin? 2.Why do RAS mutations always happen late in AML? 3.Who will relapse after VEN? nature.com/articles/s4158…
English
Vik Dhillon retweeté

Delighted to see that our work was selected as Plenary Paper in the new issue of Blood! We show that rapid clonal selection foreshadows outcome of IVO+VEN+/-AZA in AML and uncover transcriptional processes altered in resistant clones! @BloodPortfolio
ashpublications.org/blood/article-…
English

Excited to discuss our work on cathepsin protease expression and its impact on therapeutic outcomes with T-DXd in MBC while sharing insights on biomarkers that shape ADC response. Grateful for my mentor’s guidance @dradityabardia throughout this project
@OncLive #SABCS2025
OncLive.com@OncLive
It was great meeting with @AlkassisSamer, of @UCLAHealth to discuss his work evaluating cathepsin protease expression and its association with therapeutic outcomes to T-DXd in metastatic breast cancer. 🎥🔬 Stay tuned for more conference coverage as day, 2 draws to a close!
English
Vik Dhillon retweeté

A lifespan clock tells the biology of time.
Derived from millions of routine clinical records, a full‑lifecycle clock reveals insights into human development and aging, enabling early disease detection and mechanistic insights. News & Views from Nathan Price and colleagues @BuckInstitute
nature.com/articles/s4159…
English
Vik Dhillon retweeté

Proud of Jeff, our outstanding resident/future heme/onc physician, present our ex vivo drug-screening study in r/r AML at #ASH25. proof-of-con in an area with limited Rx options. Honored that he also got an ASH Abstract Achievement Award! @karmanoscancer @KarmanosHemeOnc

English
Vik Dhillon retweeté

🧵Just out in Blood @BloodPortfolio: Rapid clonal selection within early hematopoietic cell compartments presages outcome to ivosidenib combination therapy
ashpublications.org/blood/article-…
We study how resistance and durable remission emerge in IDH1-mutant AML.
English

First VA cecum in the books this week 👀
Big thanks to our @VAMinneapolis site director @DVinsard 🌟 for the elite teaching and for trusting me with some awesome EMR🔥 cases!
University of Minnesota GI Fellows@UMNGIFellows
Love to see our fellows keeping up the tradition of getting donuts for everyone when they hit their first cecum at the VA! Congratulations Dr. Itani! 💪💪💪
English
Vik Dhillon retweeté
Vik Dhillon retweeté

Natural killer cells’ functional impairment drives the immune escape of pre-malignant clones in early-stage myelodysplastic syndromes
nature.com/articles/s4146…
#MDSsm @aamdsif @MDSFoundation @ic_MDS @MDS_Hub
English
Vik Dhillon retweeté

Disease risk but not remission status determines transplant outcomes in AML: long-term outcomes of the ASAP trial @BloodPortfolio
ashpublications.org/blood/article/…
English
Vik Dhillon retweeté

(1/10) How do diverse leukemia mutations converge on the same molecular program?
In our new study @CellCellPress,the @RibackLab and @Goodell_Lab show that disparate mutations rewire shared protein networks to form nuclear condensates called coordinating bodies (C-bodies).🫧

English









