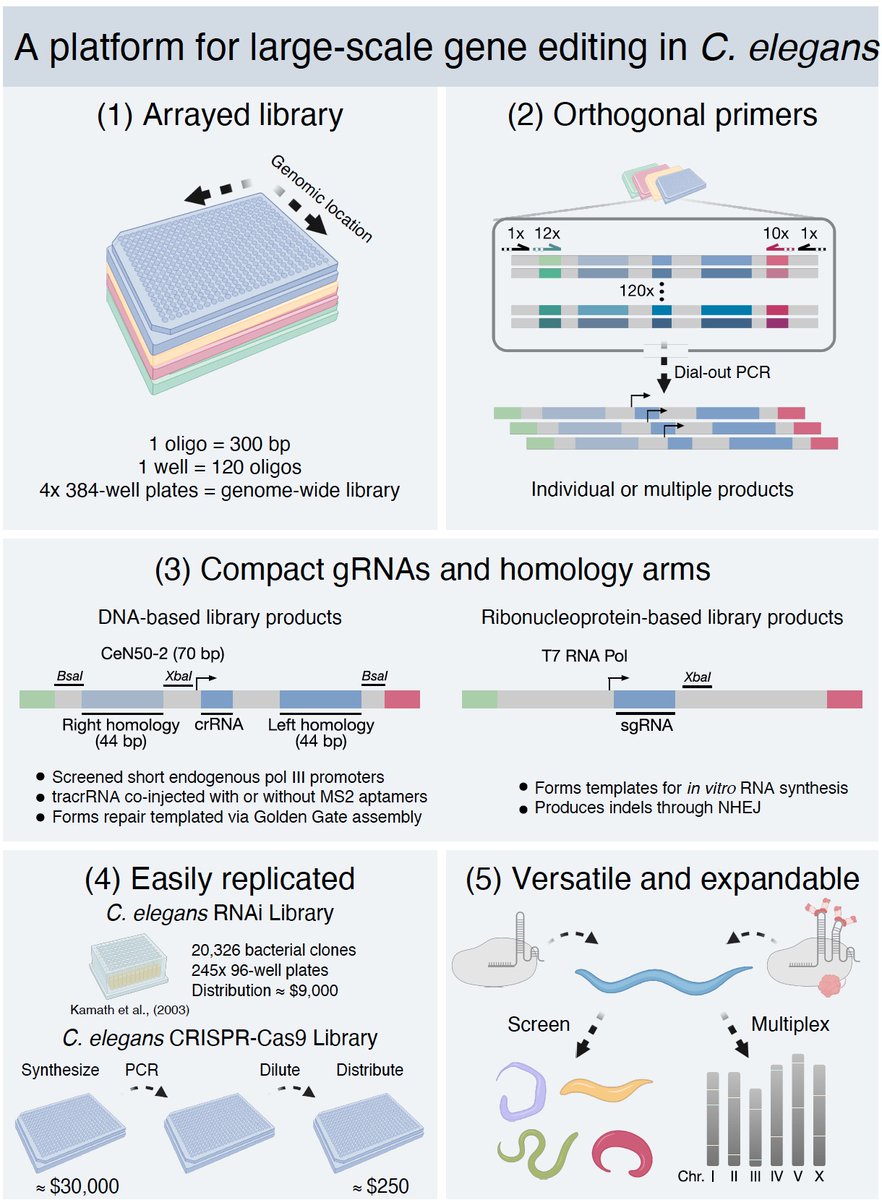
🏆Congratulations to the Top 5 onsite winners in the LEAP24 AI Oasis Hackathon! 🥇1st Place: Nuqta Genomics BioForge Project: Accelerating Genetic Engineering 🥈2nd Place: Aures DAIT Project: THE Diabetic AI Assistant 🥉3rd Place: Andromida Project: WorthyYou 🏅4th Place: RAHA Project: A voice-enabled AI for digital accessibility 🏅5th Place: Eventful Project: AI Planner They have won and joined the MVP Lab program by @ntdpsa , receiving a grant of 150,000 Saudi Riyals (40,000 US Dollars) to support and develop their winning ideas. 🚀