
MicroLive: An Image Processing Toolkit for Quantifying Live-cell Single-Molecule Microscopy biorxiv.org/content/10.110… #biorxiv_biophys
Ning Zhao
218 posts


@NingInScience
Asst. Prof. @ CU-Anschutz | K99/R00 awardee | gene regulation and protein folding | single-particle tracking | live-cell imaging | she/her

MicroLive: An Image Processing Toolkit for Quantifying Live-cell Single-Molecule Microscopy biorxiv.org/content/10.110… #biorxiv_biophys



Excited to share our latest work! We developed a new method for studying RNA localization via proximity labeling: OINC-seq! In contrast to other proximity methods, labels deposited on RNAs are read directly by sequencing without a need for biotinylation. biorxiv.org/cgi/content/sh…

Single-molecule live-cell RNA imaging with CRISPR–Csm go.nature.com/3EJPKsM


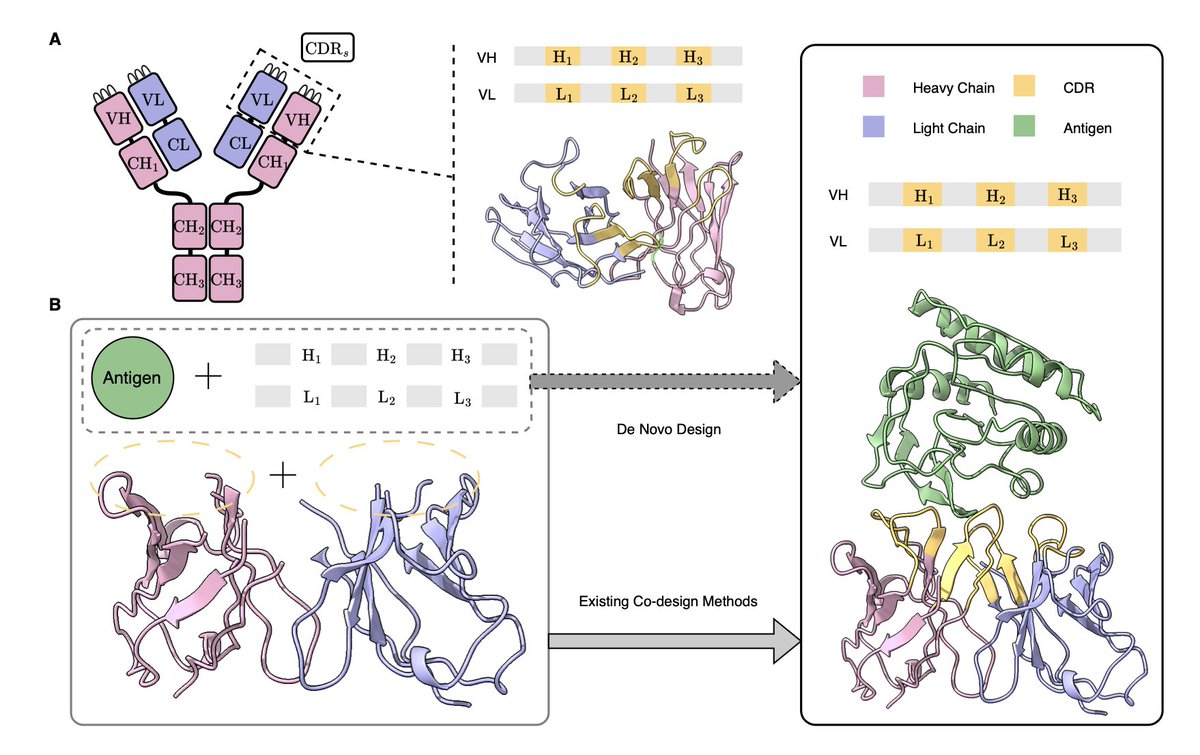






The first review paper from the Zhao lab is out today! This is a comprehensive review of not only tool development but also novel discoveries in translation. #CU_BMG #Biochem_Journal #NewPI Expanding the tagging toolbox for visualizing translation live portlandpress.com/biochemj/artic…





🧑🔬Are you thinking of doing a PostDoc? We would LOVE to “CU” here!🎉 Come to PEAKS, our Postdoc Recruitment event taking place April 7-8, 2025! 🧪 Present your Thesis work 🧑🏫 Meet our Faculty 🏫 Campus Tours 🤝 Networking Opportunities 💬 Engage with the CU Anschutz Community