
Adaptyv Bio
186 posts


@adaptyvbio
You design proteins - we run the lab. We build an automated lab that enables anyone to become a protein designer.
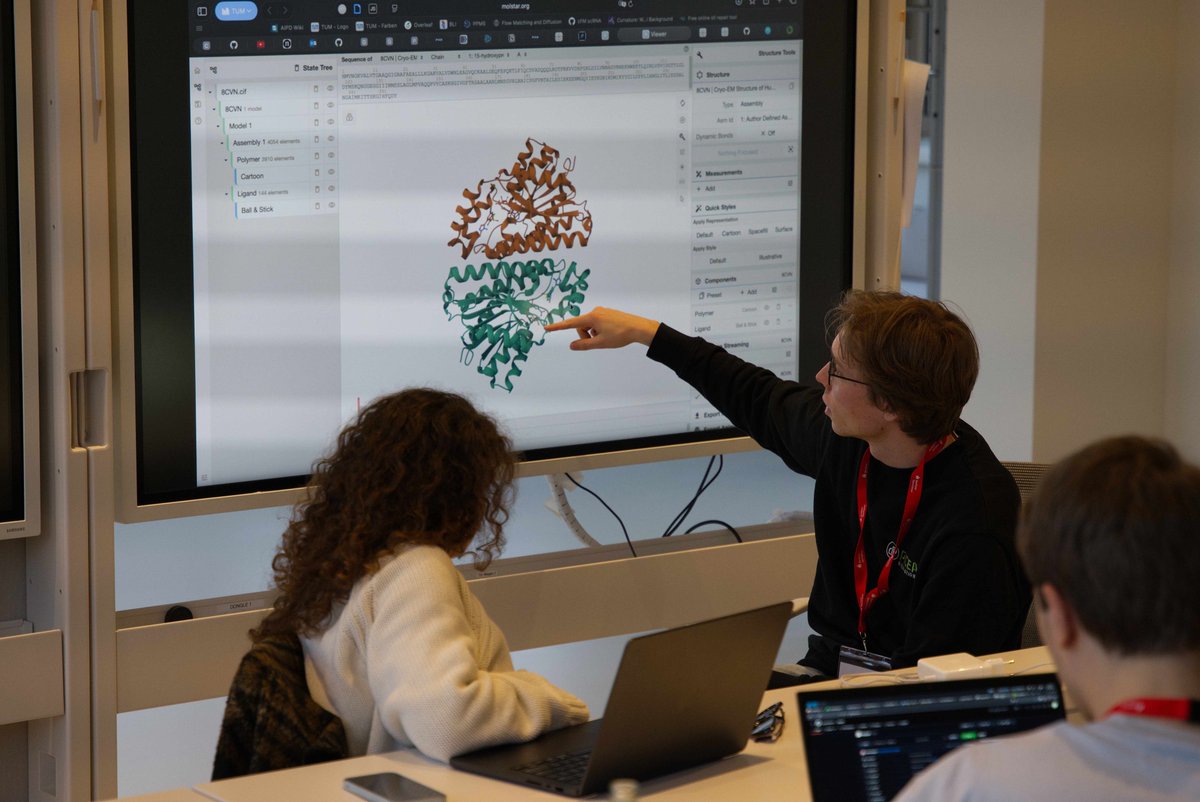















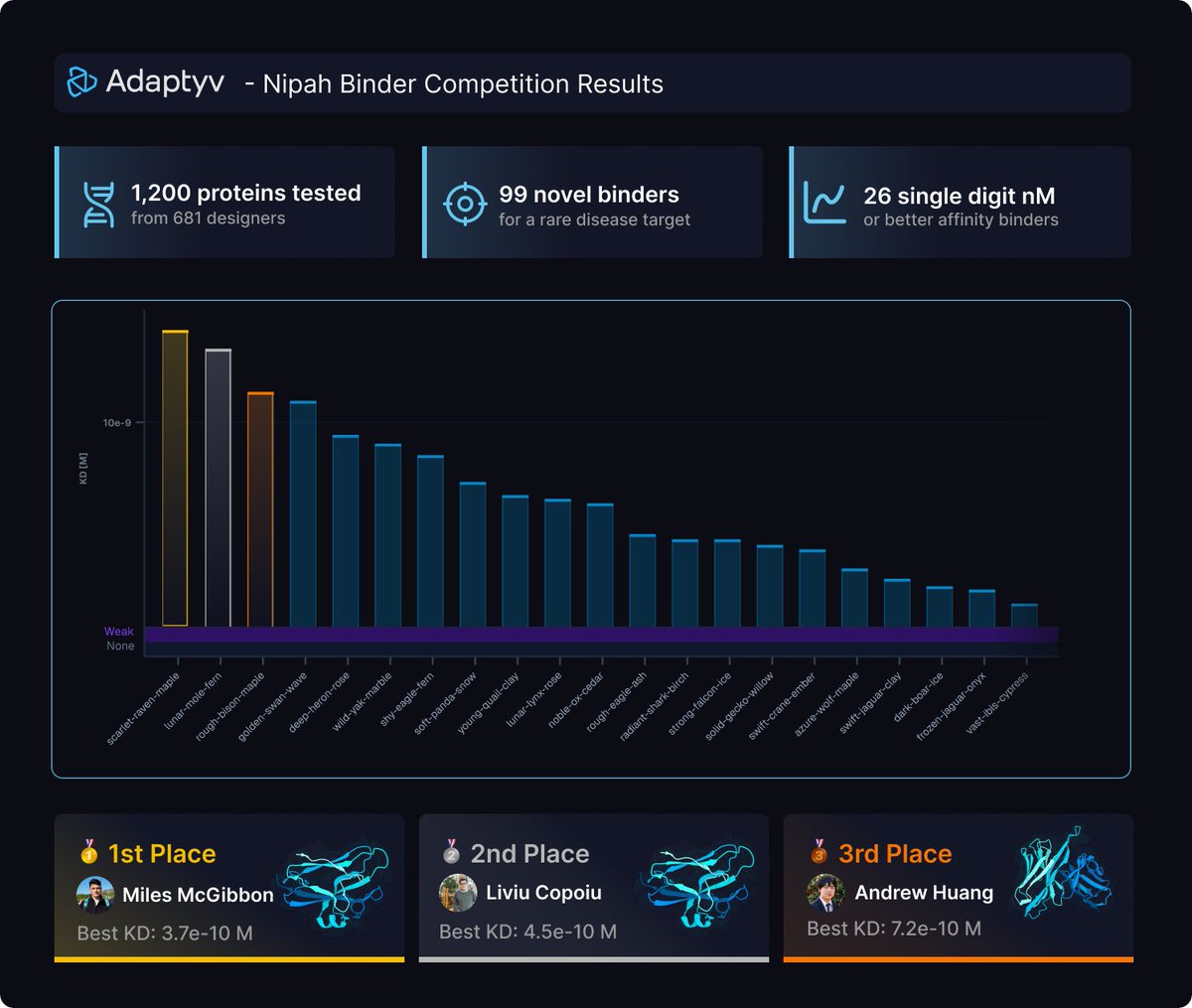
The results of the Nipah Protein Design Competition are out! 🧬 1200 proteins experimentally validated (3x more than last year) 📈 99 novel binders against the target protein (a challenging tetramer with little prior work) 💪 26 single digit nM or better binders, with the best ones at single-digit picomolar affinity! All data now available open-source on Proteinbase! Let's take a look at the results ⬇️

The results of the Nipah Protein Design Competition are out! 🧬 1200 proteins experimentally validated (3x more than last year) 📈 99 novel binders against the target protein (a challenging tetramer with little prior work) 💪 26 single digit nM or better binders, with the best ones at single-digit picomolar affinity! All data now available open-source on Proteinbase! Let's take a look at the results ⬇️


