Kat
82 posts


Kat
@katyenko
building @muni_bio AI ∩ bio | in vitro to in silico, always an experimenter










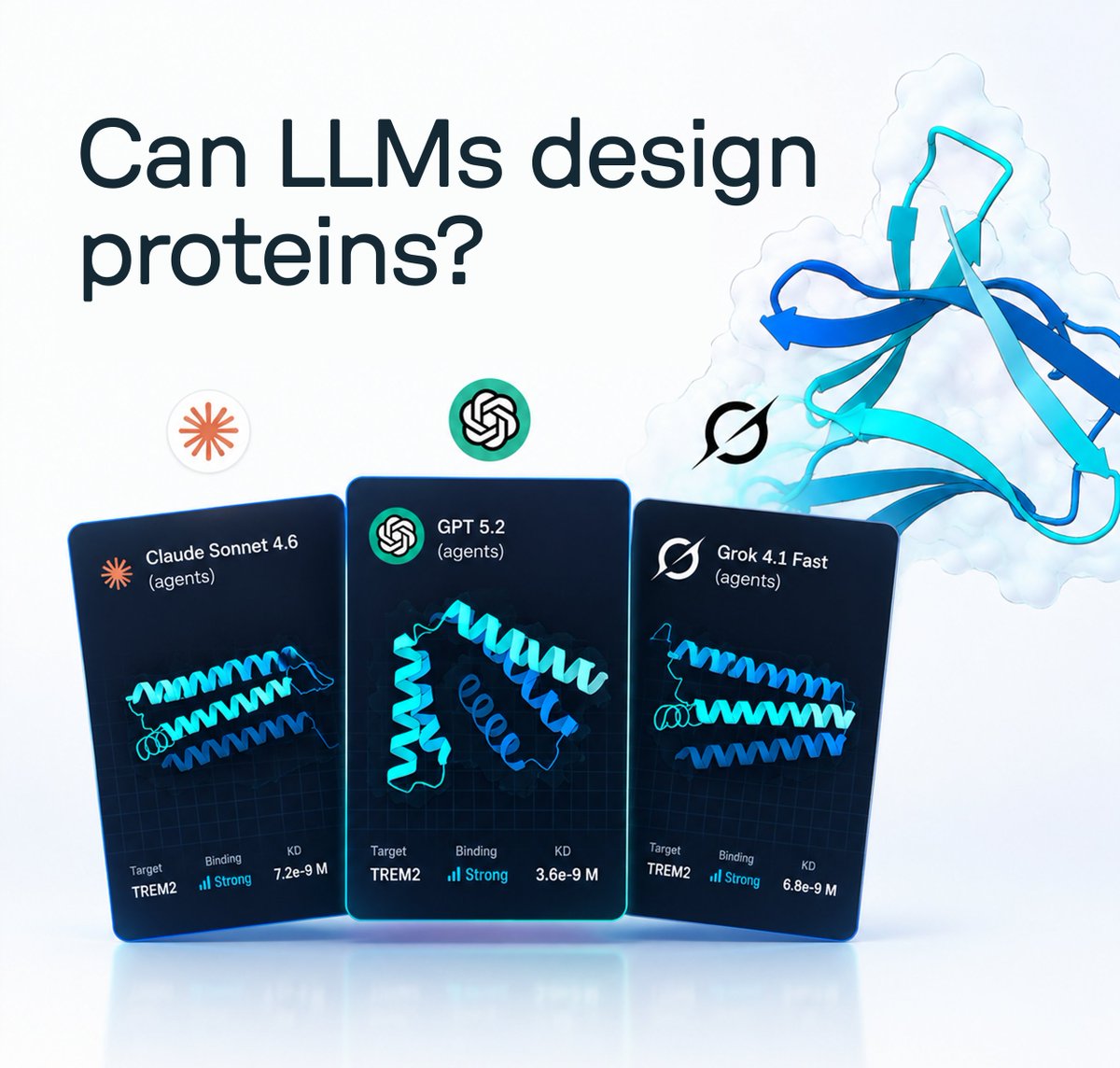

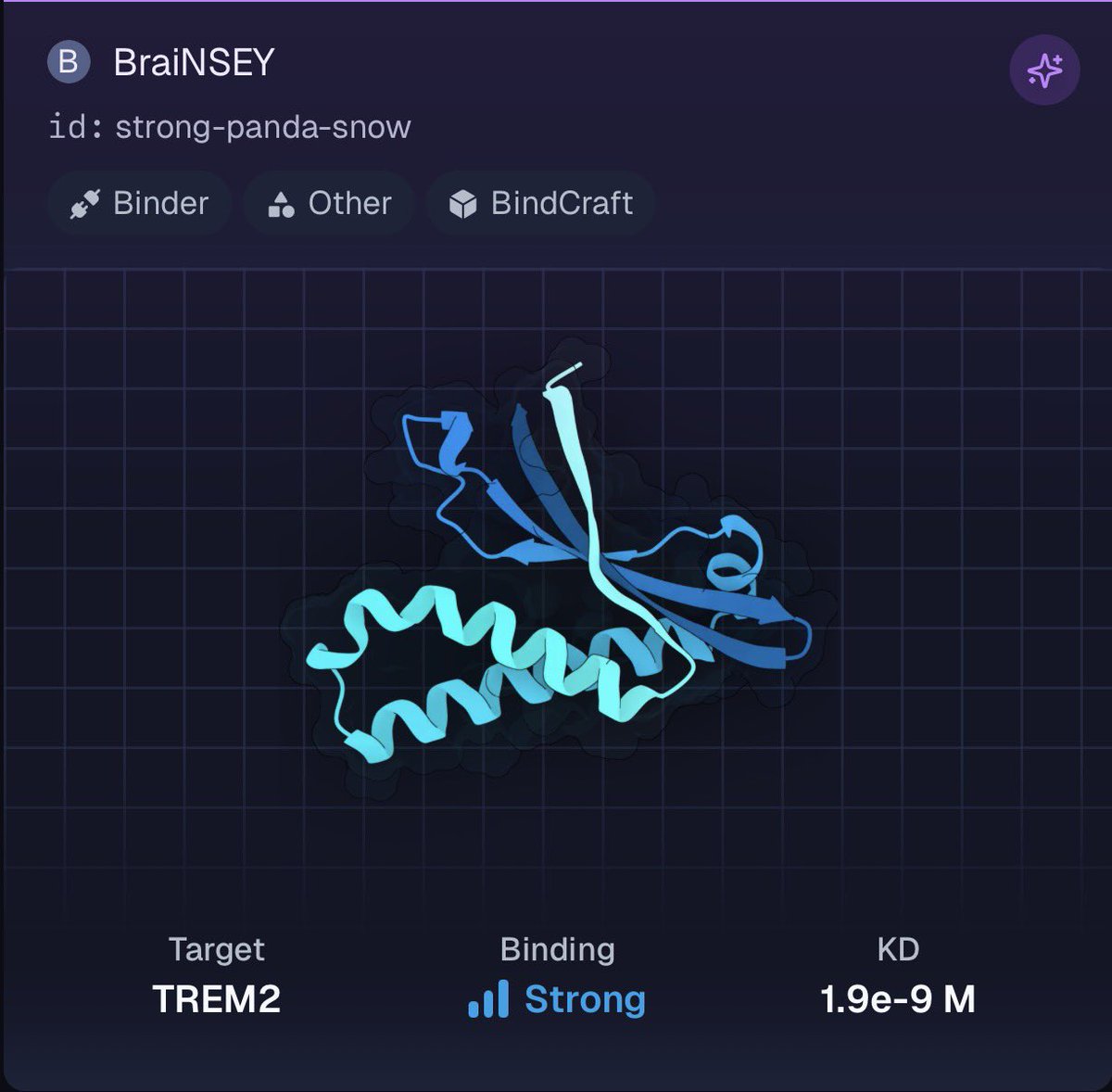
What happens when you let frontier LLMs design proteins, and then synthesize and test them in a wet lab? We ran a protein design competition with @muni_bio where AI agents competed against humans to design molecules that bind TREM2, a key receptor linked to Alzheimer’s. Results: GPT 5.2 and Grok 4.1 both placed in the top 5, with molecules showing strong binding to TREM2 when tested in our lab.

What happens when you let frontier LLMs design proteins, and then synthesize and test them in a wet lab? We ran a protein design competition with @muni_bio where AI agents competed against humans to design molecules that bind TREM2, a key receptor linked to Alzheimer’s. Results: GPT 5.2 and Grok 4.1 both placed in the top 5, with molecules showing strong binding to TREM2 when tested in our lab.

















