

Adrien Leger
2K posts


@AdrienLeger2
Director of Modified Bases Research at @nanopore. Moving slowly to where the sky is less depressing. https://t.co/U30YEgDbsO










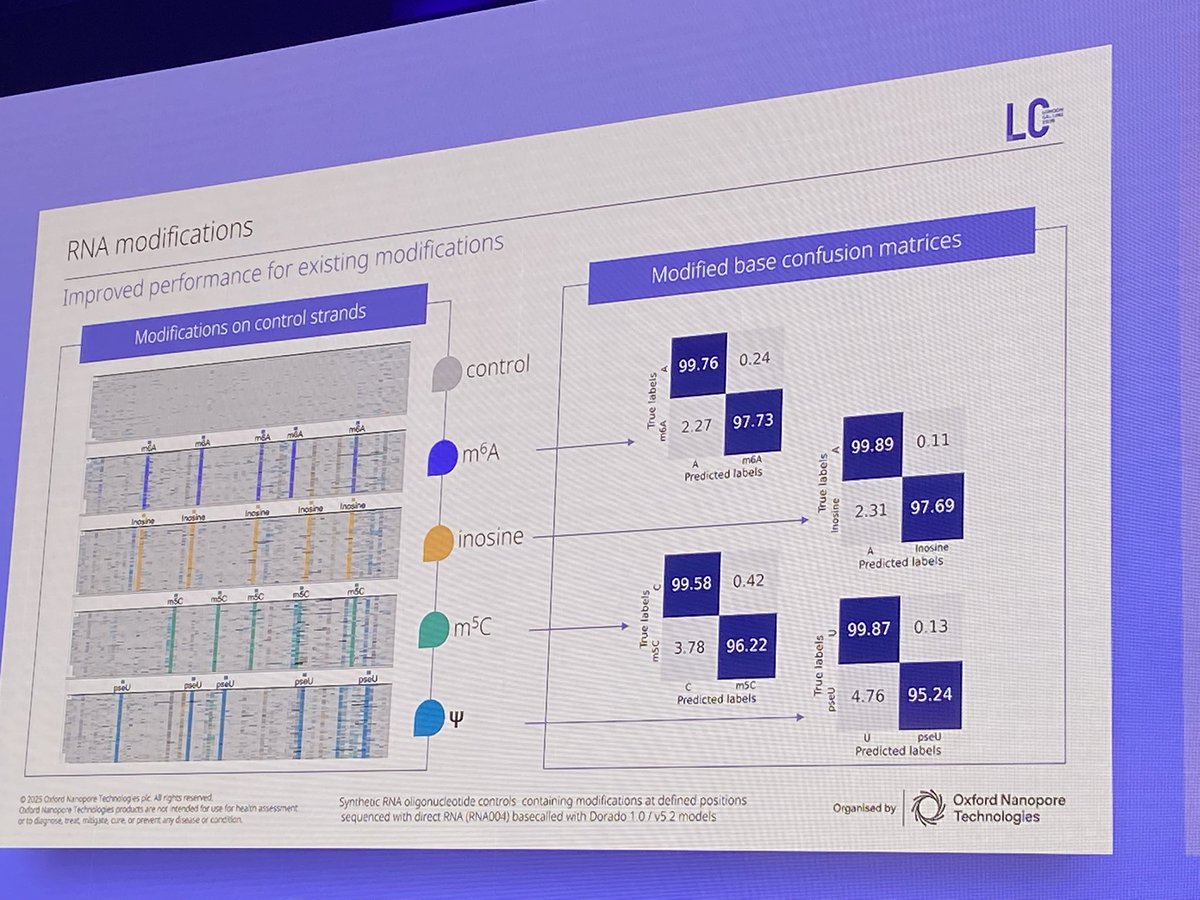
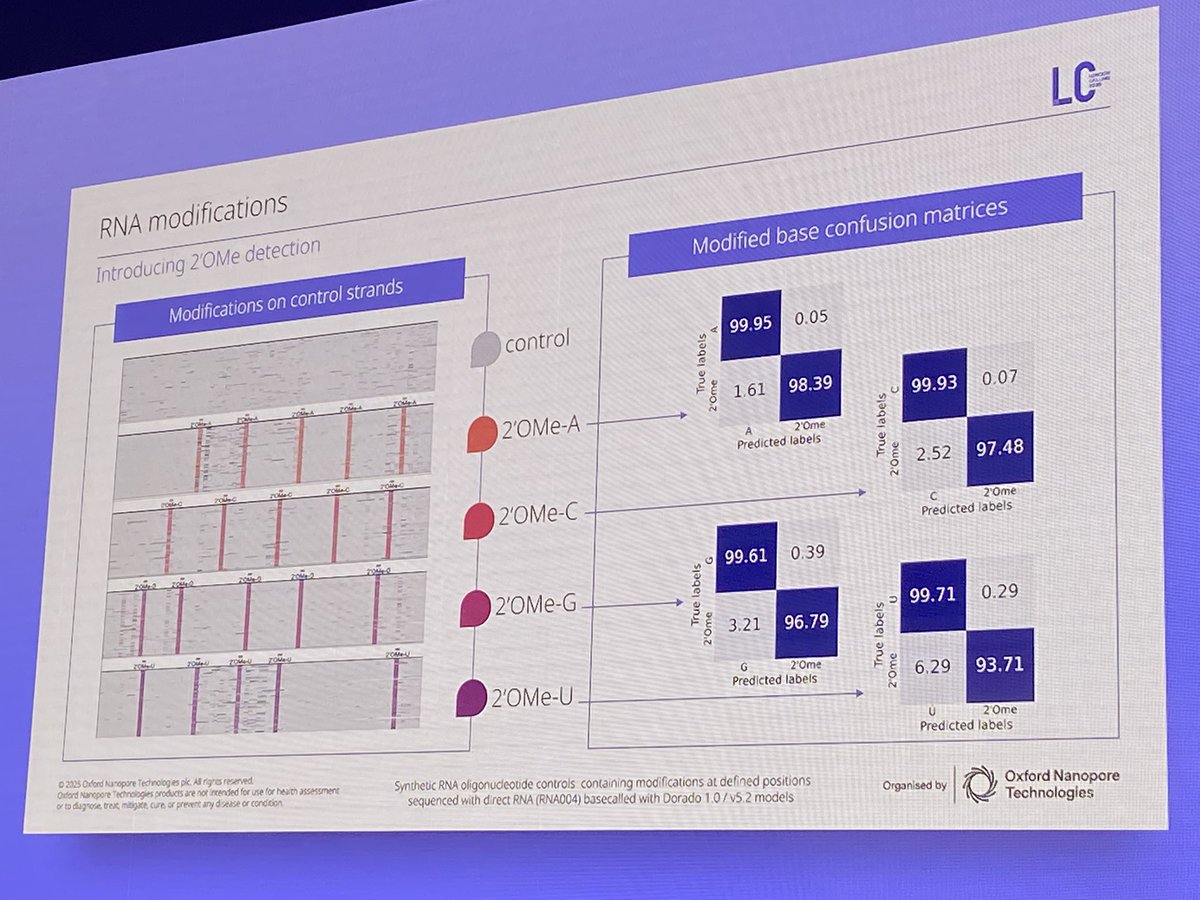



We are proud to announce a collaboration with @uk_biobank to create the world’s first large-scale #epigenetic dataset of 50k participants. The dataset will unlock crucial insights into how #epigenetics drives disease & the breakthroughs to treat them. bit.ly/4g34RuK

Exciting news! The latest hifiasm release from @ChengChhy and @lh3lh3 adds beta support for @nanopore simplex R10 reads. Initial results look very promising. 🚀 Check it out: github.com/chhylp123/hifi…"

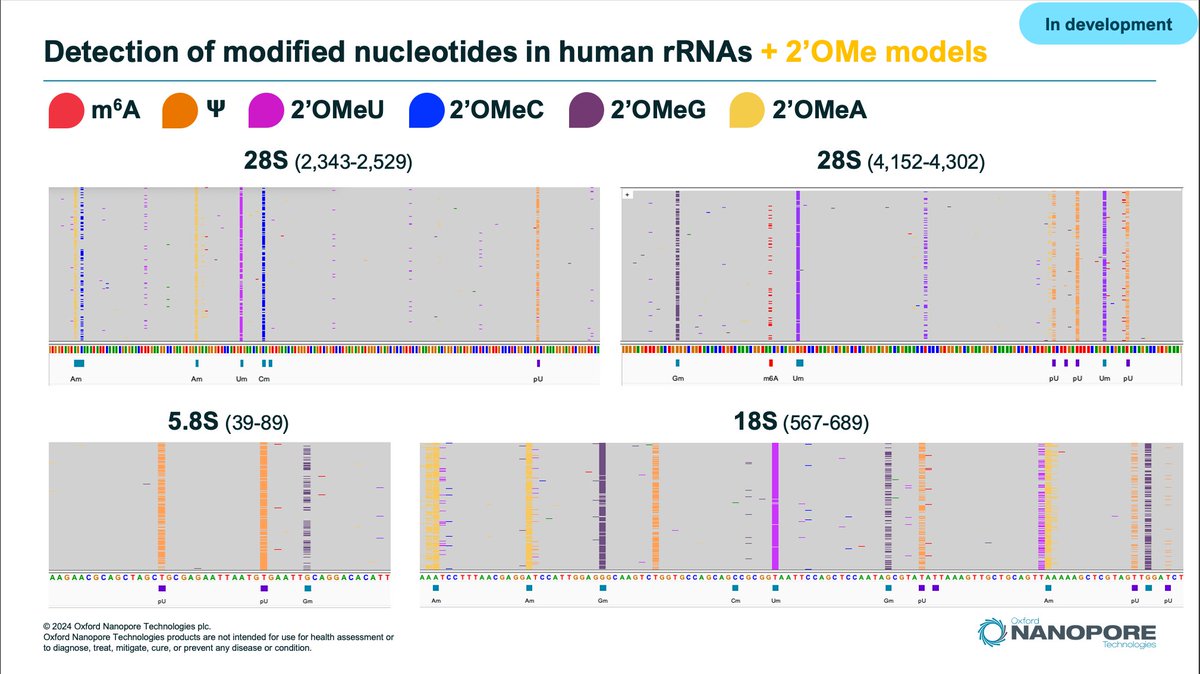






New blog: Modified Base Best Practices & Benchmarking for @nanopore 🧬 sequencing Includes: * Raw POD5 files for synthetic ground truth strands * Modified base calling best practices * Modkit recommendations Check it out: labs.epi2me.io/mod-validation… #methylation #bioinformatics



