
Nathan 🧬
291 posts


@nathanbaggers
Sequencing Chemistry Research Scientist @nanopore Views are my own







We are proud to announce a collaboration with @uk_biobank to create the world’s first large-scale #epigenetic dataset of 50k participants. The dataset will unlock crucial insights into how #epigenetics drives disease & the breakthroughs to treat them. bit.ly/4g34RuK



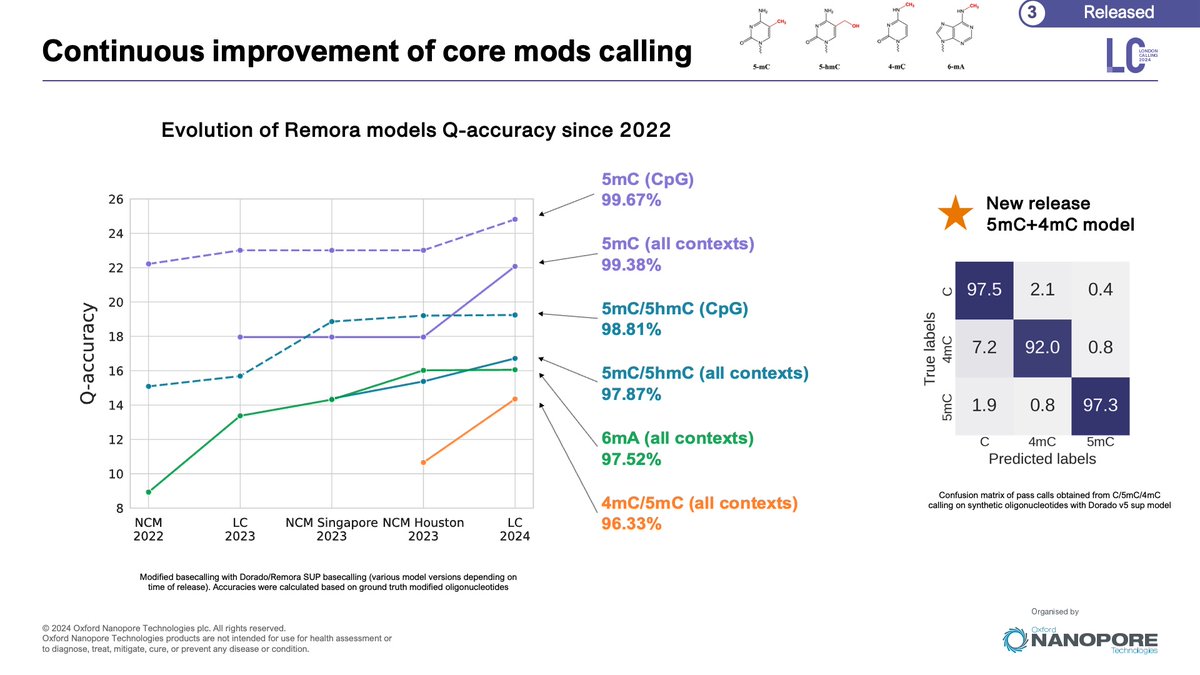
New blog: Modified Base Best Practices & Benchmarking for @nanopore 🧬 sequencing Includes: * Raw POD5 files for synthetic ground truth strands * Modified base calling best practices * Modkit recommendations Check it out: labs.epi2me.io/mod-validation… #methylation #bioinformatics

.@nanopore today announced that @RosemaryDokos has been appointed Chief Product & Marketing Officer & Lakmal Jayasinghe as Chief Scientific Officer, effective immediately. Rosemary & Lakmal succeeded Clive Brown as Chief Officer of Technology, Innovation, & Products. Read more: bit.ly/3O4ulvQ



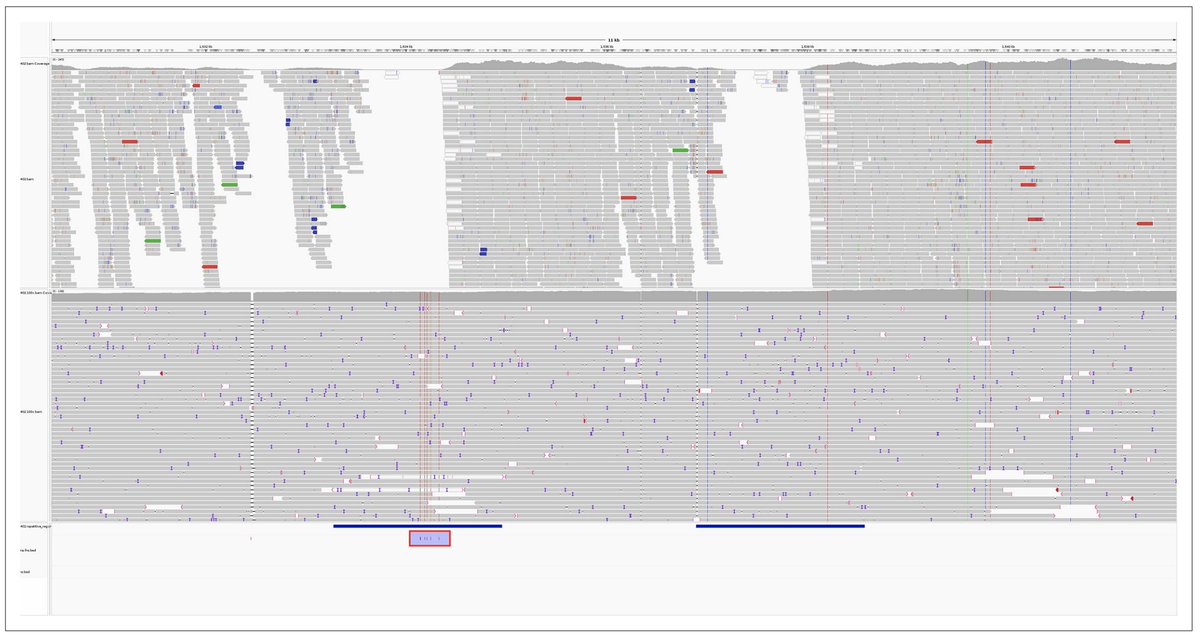

“This important study shows how a combination of the latest generation of @nanopore long reads with state-of-the art variant callers enables bacterial variant discovery at an accuracy that matches or exceeds the current "gold standard" with short reads” elifesciences.org/articles/98300
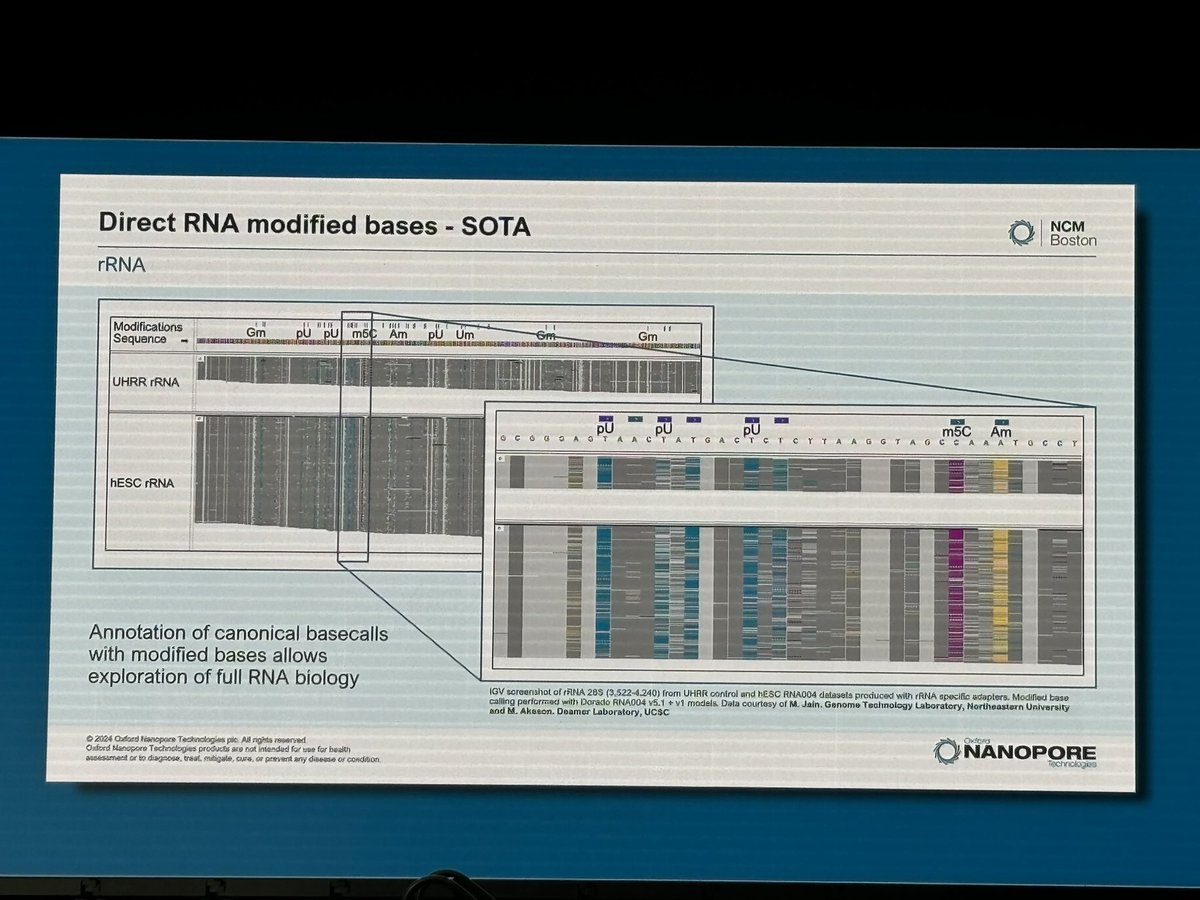









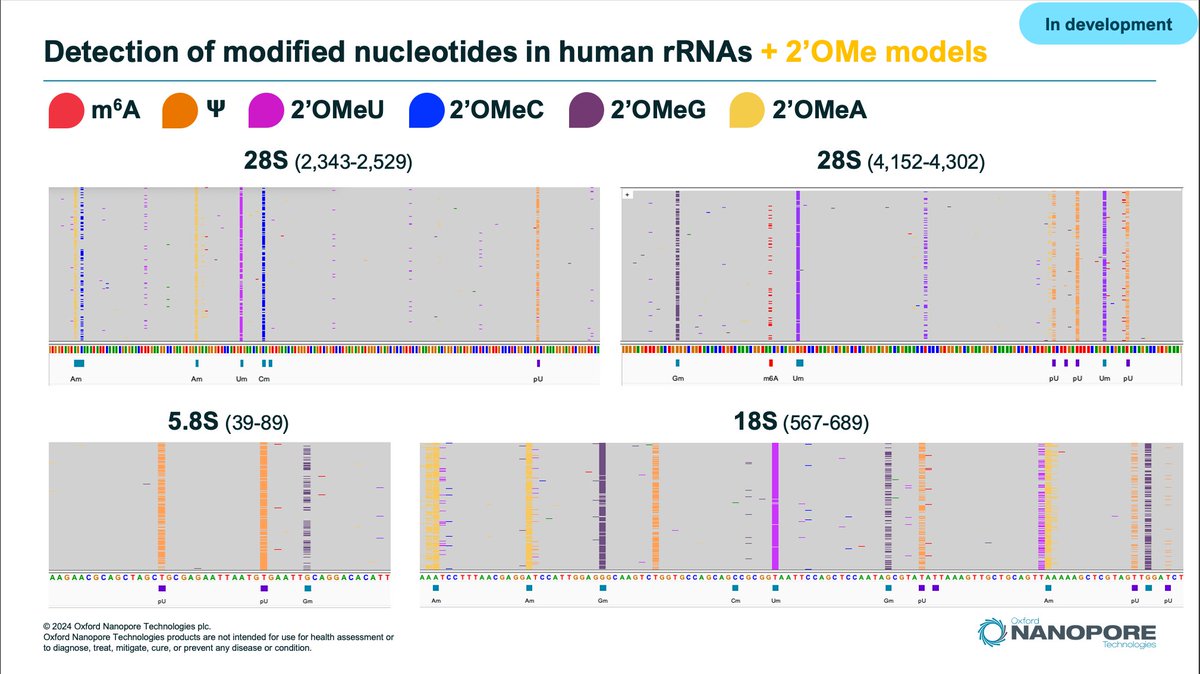
Dorado v0.8.0 #ncm24 release! - v5.0 DNA transformer model performance improved for A100/H100 via kernel fusing, quantisation and leveraging i8 tensor cores. - v5.1 RNA models with updated mod calling for m6A, pseU, and brand new calling of Inosine and m5C. - Improved dorado correct performance, stability and flexibility. - Introduces per-barcode configuration for poly(A) estimation with interrupted tails. #nanoporeconf github.com/nanoporetech/d…




