

Alisandra Denton (@[email protected])
151 posts


@AlisandraDenton
Senior Machine Learning Scientist @ Valence labs Bringing the power of deep learning to understanding our cells and DNA. https://t.co/9NVX6q3fFM…










🛡️ New Certified Dataset and Benchmark! Did you know that high-dimensional data like cell images (i.e. phenomics) can help us understand the relationship between compounds and genes? Now you’ll have a chance to work with a phenomics dataset and benchmark to predict which genes a given compound interacts with! RxRx3-core is the latest certified dataset from @RecursionPharma that includes labeled images of 735 genetic knockouts and 1,674 small-molecule perturbations. The related benchmark evaluates the zero-shot prediction of compound-gene activity. If you’re at NeurIPS, come talk to the Polaris and @RecursionPharma teams! We’ll be at the conference and workshops over the weekend. Explore the dataset: polarishub.io/datasets/recur… Explore the benchmark: polarishub.io/benchmarks/rec… Chime in on the certification process: github.com/polaris-hub/po…
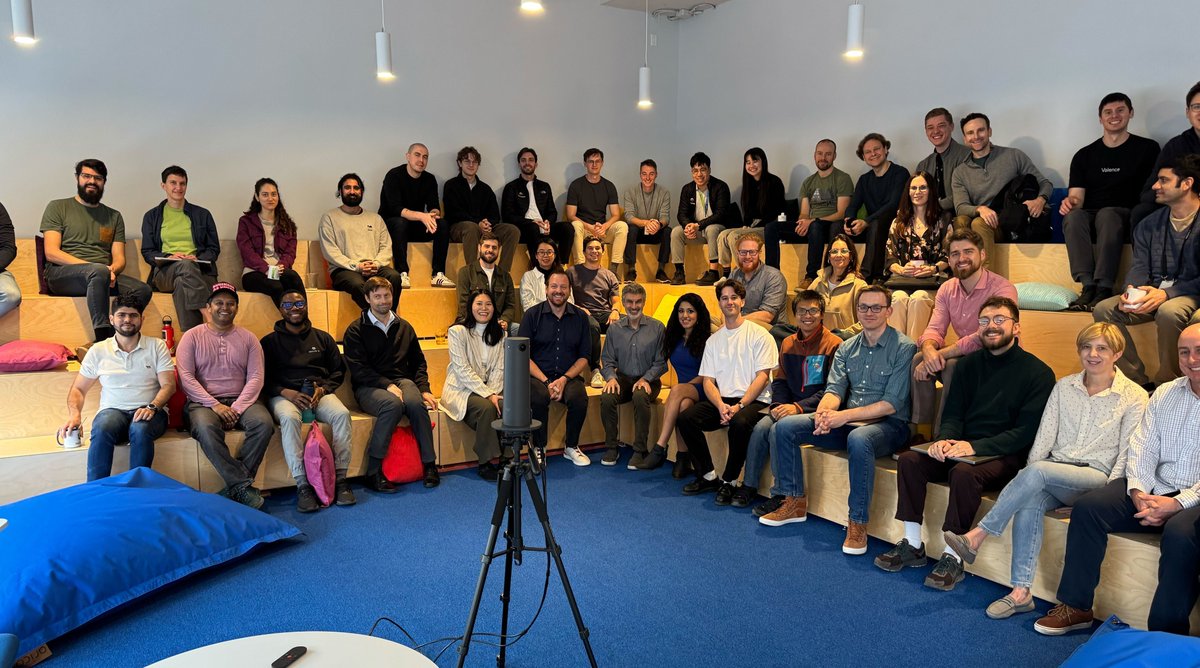















Next stop: LONDON! 🇬🇧 We’re expanding our operations to Europe, at the epicenter of the rapidly growing #TechBio sector in #London. Learn more at bit.ly/3wOCSO2 and check out our new open positions at recursion.com/careers $RXRX #biotech #ai #ml

Next stop: LONDON! 🇬🇧 We’re expanding our operations to Europe, at the epicenter of the rapidly growing #TechBio sector in #London. Learn more at bit.ly/3wOCSO2 and check out our new open positions at recursion.com/careers $RXRX #biotech #ai #ml

Next stop: LONDON! 🇬🇧 We’re expanding our operations to Europe, at the epicenter of the rapidly growing #TechBio sector in #London. Learn more at bit.ly/3wOCSO2 and check out our new open positions at recursion.com/careers $RXRX #biotech #ai #ml