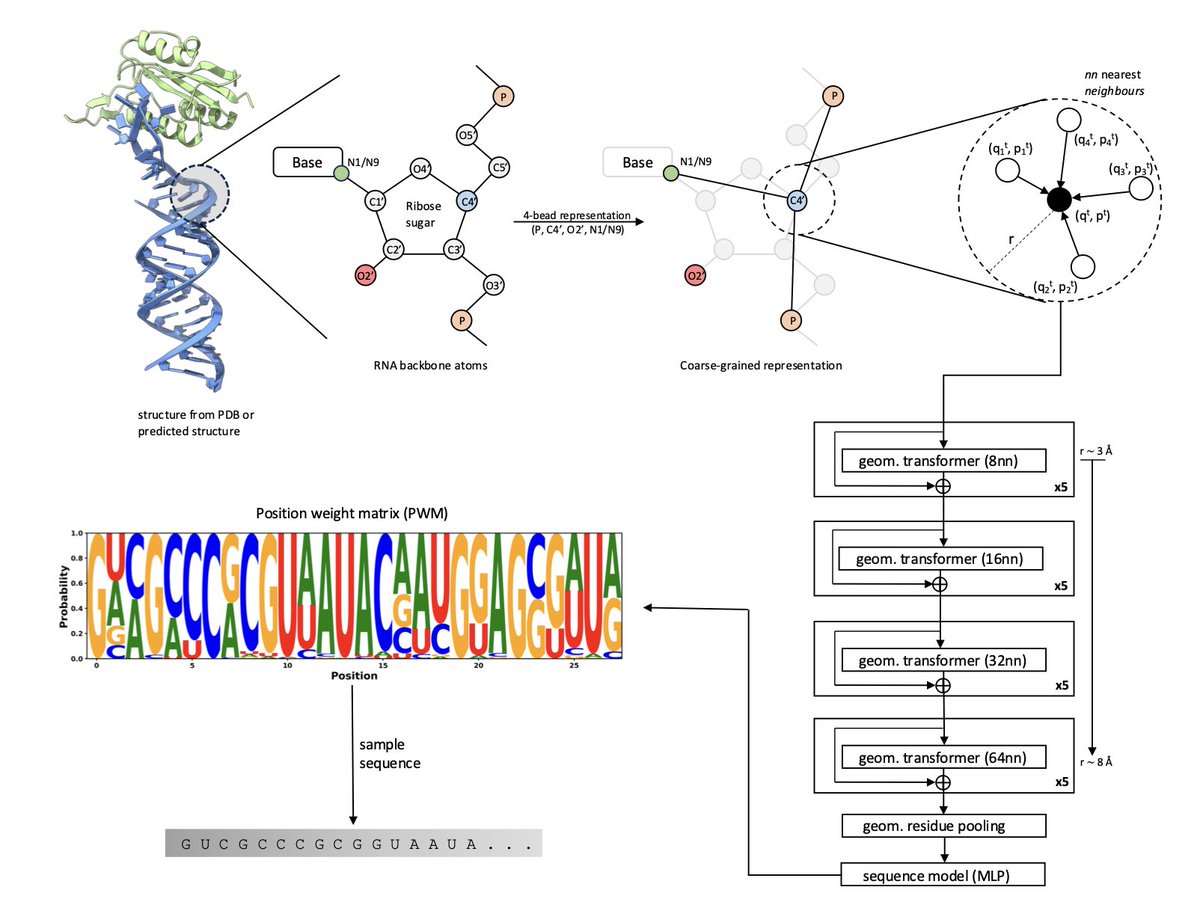
Presenting RISoTTo, a molecular context-aware model for RNA sequence design, building on our work with PeSTo (for protein binding interface prediction) and CARBonAra (for protein design). 📄Preprint: biorxiv.org/content/10.110… 💻Code & more coming soon! #RNAbiology #AI4Science