Christoph Hahn
1.4K posts

Christoph Hahn
@C__Hahn
Biology, Bioinformatics | husband & father | curious, fascinated, frustrated, exhausted, excited, hungry, surprised ..
Katılım Ekim 2014
193 Takip Edilen246 Takipçiler
Christoph Hahn retweetledi

Our preprint is out! We hacked the @nanopore sequencer to read amino acids and PTMs along protein strands. This opens up the possibility for barcode sequencing at the protein level for highly multiplexed assays, PTM monitoring, and protein identification!
biorxiv.org/content/10.110…
English
Christoph Hahn retweetledi

Yesterday we released our preprint showing how a @nanopore array morphs into a protein sequencer... Achieving LONG reads (hundreds of amino acids) and the capability to re-read individual molecules multiple times, all thanks to a protein-processive molecular motor.
Keisuke Motone / 元根 啓佑@KeisukeMotone
Our preprint is out! We hacked the @nanopore sequencer to read amino acids and PTMs along protein strands. This opens up the possibility for barcode sequencing at the protein level for highly multiplexed assays, PTM monitoring, and protein identification! biorxiv.org/content/10.110…
English
Christoph Hahn retweetledi

Evolutionary dynamics of whole-body regeneration across planarian flatworms nature.com/articles/s4155…

English

Embracing the command line: my unexpected career in computational biology nature.com/articles/d4158…
English
Christoph Hahn retweetledi

Great news from COPO!
Reliably brokering sample #metadata is an essential step of the reference #genome production workflow.
Congratulations on this impressive milestone 🎉
@BioGenEurope @EarlhamInst
English
Christoph Hahn retweetledi

Structural #genome annotation with BRAKER & TSEBRA - workshop will take place on the 21st Nov. from 9am to 3pm, registration is free but capacity is limited to 35 participants - so hurry! cognitoforms.com/ERGAEuropeanRe… #training #bioinformatics #genomics #biodiversity @erga_biodiv ERGA
English
Christoph Hahn retweetledi

Thrilled that our work “compleasm: a faster and more accurate reimplementation of BUSCO” is now published in Bioinformatics! 🎉 Huge thanks to our amazing lab @lh3lh3 and to the reviewers’ valuable comments. 🙏 Journal paper: academic.oup.com/bioinformatics…
English
Christoph Hahn retweetledi

In a post #AlphaFold world, we can use protein structures in ways we never could before. Can we build phylogenies with them? Are they any good? Yes! Foldtree (biorxiv.org/content/10.110…) surpasses traditional sequence-based methods, even for closely related proteins.👇

English

Indeed, thanks @philippresl, @EcoEvoGraz, @UniGraz. Check it out - even if you're not interested in #phylogenetics. It's about democratizing complex computational analyses and #reproducibiltiy through #scalability and #portability (#containers)..
Philipp Resl@philippresl
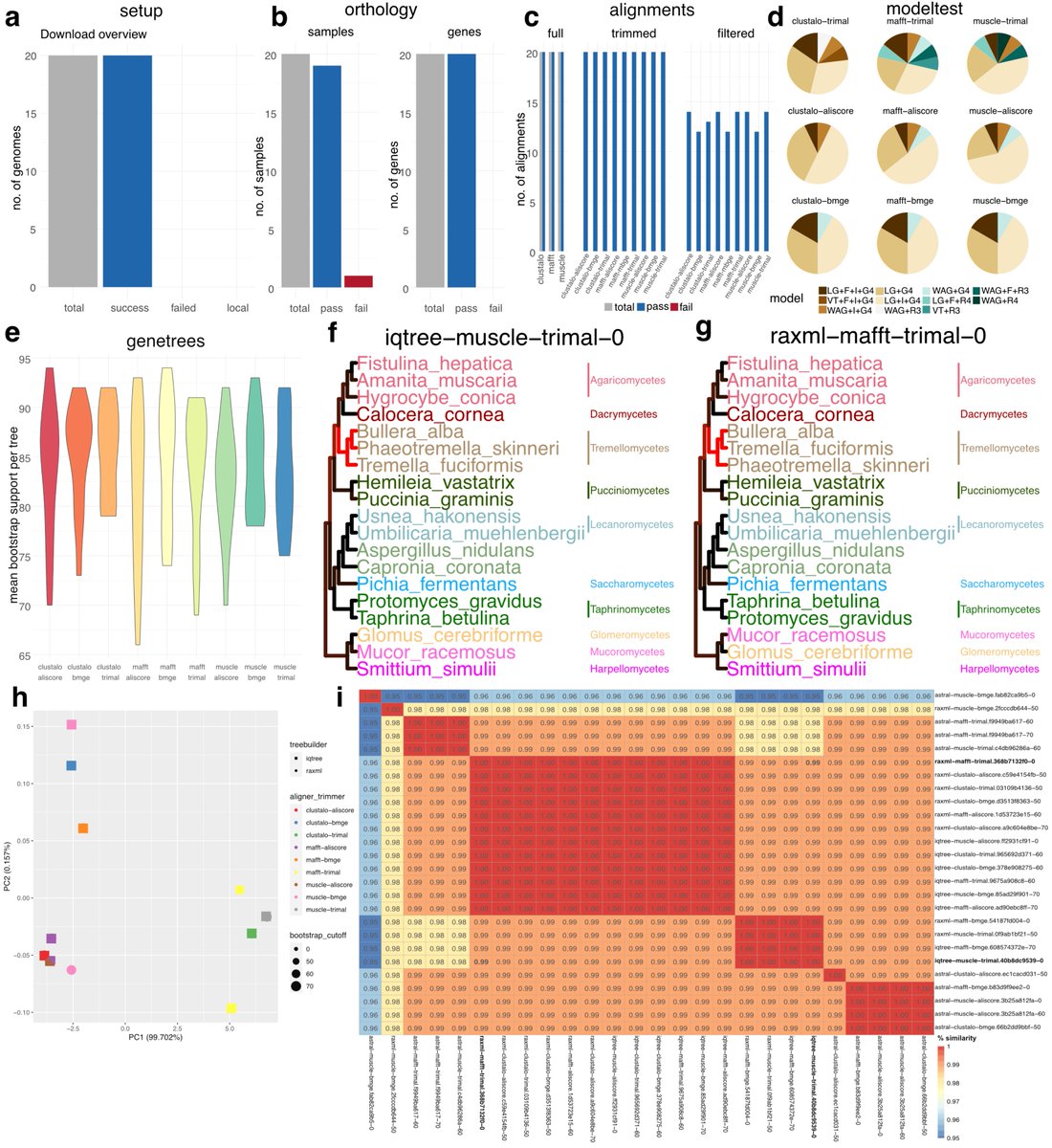
Interested in #phylogenetics using whole #genome/#transcriptome/#proteome data? @C__Hahn and I are very happy to share phylociraptor with you: It facilitates explorative and reproducible phylogenomic analyses of thousands of genes and genomes. Preprint: tinyurl.com/yc4f7v6d
English
Christoph Hahn retweetledi
Christoph Hahn retweetledi

Interested in #phylogenetics using whole #genome/#transcriptome/#proteome data? @C__Hahn and I are very happy to share phylociraptor with you: It facilitates explorative and reproducible phylogenomic analyses of thousands of genes and genomes. Preprint: tinyurl.com/yc4f7v6d
English
Christoph Hahn retweetledi

Phylociraptor - A unified computational framework for reproducible phylogenomic inference biorxiv.org/cgi/content/sh… #bioRxiv
Română
Christoph Hahn retweetledi

Phylociraptor - A unified computational framework for reproducible phylogenomic inference biorxiv.org/cgi/content/sh… #biorxiv_bioinfo
Română
Christoph Hahn retweetledi

Our CARD pilot paper with @KimberleyBill10 @cornelisblauw @BenedictPaten and many others is online! We establish a wet + dry lab protocol for single nanopore flow cell sequencing, which generates state-of-the-art SNP and SV calls + assemblies. rdcu.be/dl901
English
Christoph Hahn retweetledi

Snakemake has many technical benefits over Nextflow for writing bioinformatics workflows.
But there is no platform that supports the language.
Today we are releasing native support for Snakemake on the LatchBio Cloud.
blog.latch.bio/p/native-snake…
English
Christoph Hahn retweetledi

The small variant calling benchmarking initiative of @NGSCN1 is published. Thanks a lot to all the contributors and @dfg_public for the funding. (1/x)
English

Visual opsin gene expression evolution in the adaptive radiation of cichlid fishes of Lake Tanganyika | Science Advances science.org/doi/10.1126/sc…
English
Christoph Hahn retweetledi

Assessment of metagenomic workflows using a newly constructed human gut microbiome mock community. #Metagenomics #MicrobiomeMockCommunity #DNAresearch academic.oup.com/dnaresearch/ar…
English
Christoph Hahn retweetledi

How do you stress the importance of replicates to improve the accuracy of #eDNA sampling?
Use pizza of course 🍕🍕🍕
A great talk by @N_P_Griffiths
#fishimp2023
English