Sabitlenmiş Tweet
Kenny Workman
1.6K posts

Kenny Workman
@kenbwork
cto @latchbio, data infrastructure for biology
Katılım Mayıs 2020
1.3K Takip Edilen7K Takipçiler

@gchies1 Agree with this in practice but opted to collapse this complexity to make accessible to broader audience
English

@kenbwork Good post but mentioning complex techniques aas “you just do a perturb-Seq” seems to me a bit dismissive of the complexity each techniques involved. Beside their cost. There is a hidden cost of complexity I.e. how much trial/error and experience it takes to ace the experiment
English

Agents calling tools now generate more revenue on Latch than scientists clicking buttons. From native deployments but also harnesses like Claude Code, Codex, Cursor across Pharma, biotech, academic labs. Product roadmap increasingly concerned with exposing compute intensive operations and experimental context to agents while designing new ways for humans to steer. Tool calls can dispatch a TB of data over a dozen computers. Errors in eg. parameter selection, infra failures, tool selection more expensive in time/cost than familiar agentic domains. Very early and exciting product space.
English

Models still struggle with biology-specific analysis details. Many remaining evals require knowledge of experimental design, kit- or instrument-specific analysis, or tacit scientific knowledge. Examples include:
- Aggregating to the correct statistical unit before testing, such as inappropriately treating cells as statistical replicates and inflating power
- Applying whole-transcriptome scRNA-seq defaults to targeted 30–400 gene panels, such as BD Rhapsody
- Accurately recognizing and removing depth or batch confounds, such as PC1 absorbing library size
- Integrating data from multiple platforms, such as DNA + RNA on Tapestri or protein + mRNA
English

Trajectory review reveals improvement in exploratory behavior:
- Thoroughly investigating data and experimental context before choosing an analysis algorithm or tool
- Self-verifying after intermediate steps and before submitting answers
- Early signs of assay-aware analysis, such as QC thresholds specific to kit or instrument type instead of generic tutorial defaults. This largely explains the jump in Opus 4.7 accuracy.
English

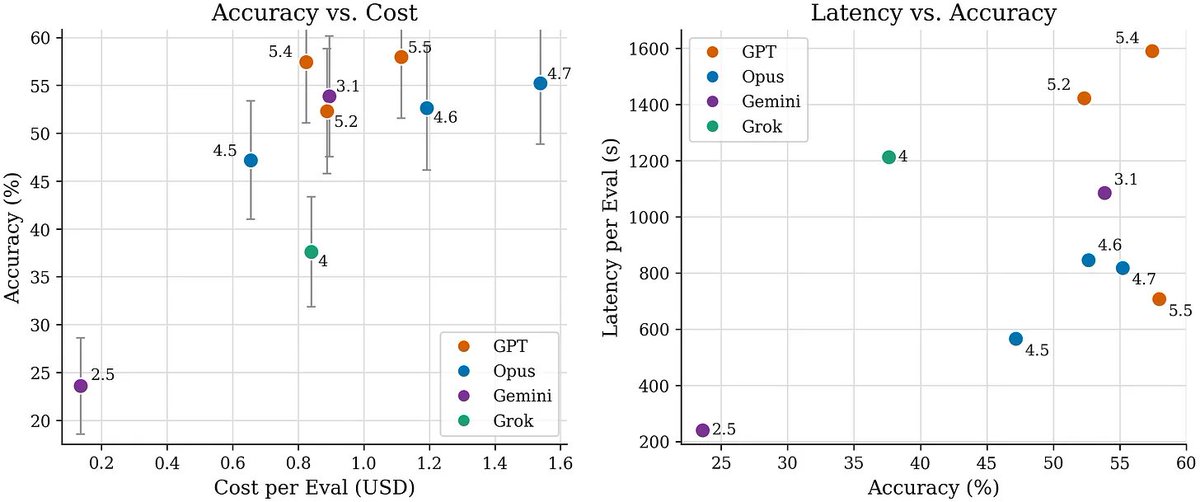
Are frontier models improving at single cell biology?
We revised scBench with additional rounds of scientist review and generated new benchmark data for Opus 4.7, GPT 5.5, and Gemini 3.1.
On accuracy, Gemini 3.1 roughly doubles, Opus 4.7 shows modest improvement, and GPT 5.5 shows little to no improvement.
On latency, GPT 5.5 improves substantially, Opus 4.7 shows little change, and Gemini 3.1 worsens by ~5x.
English
Kenny Workman retweetledi

This was from work released late December and model capabilities have improved since then. See benchmarks.bio.
As models climb procedural data analysis skill, longer horizon tasks (eg. what a biologist might publish in the results section of a paper) become soluble. More to share on efforts there soon.
English

Some relevant links:
YouTube: youtube.com/watch?v=PeTn4x…
Slides: docs.google.com/presentation/d…
Preprint: arxiv.org/abs/2512.21907
Thank you to @arshianazar @AgaRosegirl + @BerkeleyML for having me.

YouTube
English

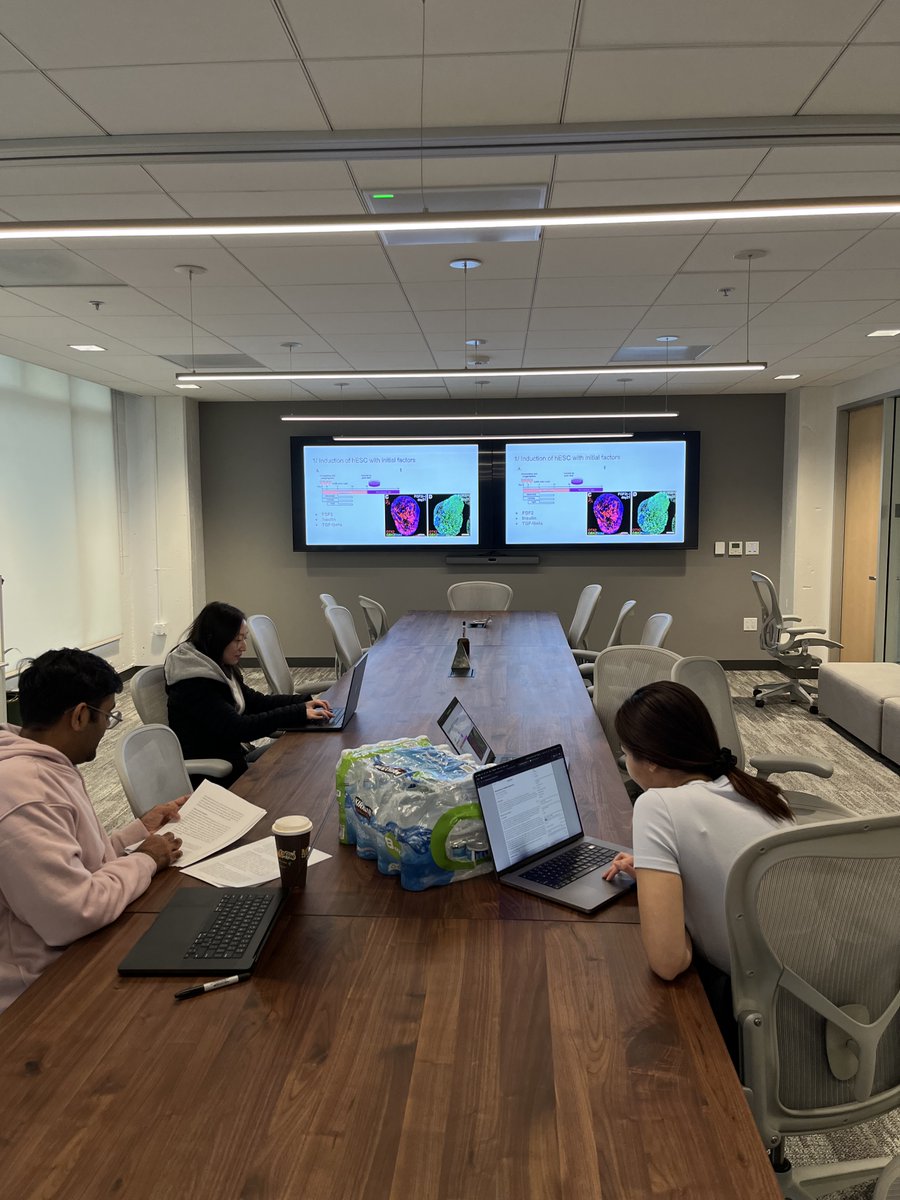
Technical talks on engineering challenges with AI and single-cell data in Mission Bay, SF, next Thursday.
Material covering emerging analysis methods for new kits, benchmarks and evaluations for frontier models, and practical AI for drug screening.
Harihara Muralidharan — Technical Staff @ LatchBio
Valentine Svensson — Principal Computational Biology Scientist @ Tahoe Therapeutics
Mikaela Koutrouli — Core Developer @ scverse
Zhen Yang — Technical Staff @ LatchBio
Link below:

English

The platforms these assay providers have built on top of Latch are incredible. Each reflects months of customer obsession and scientific depth. Some of my favorites:
- vizgen.latch.bio
- takarabio.latch.bio
- atlasxomics.latch.bio
- brokenstring.latch.bio
English

Latch almost died two years ago. Selling data infrastructure direct to pharma was not leading to sustainable growth. We found a group of fast moving customers closer to the source of data generation: teams engineering kits + machines for single cell, spatial and proteomics assays. They brand and distribute Latch, focusing on difficult scientific problems specific to how each assay works, while we focus on keeping infra performant as data throughput continues to ride exponential. Today our team is nearly 50 scientists and engineers with one dedicated sales engineer. Have learned an incredible amount about the internals and nuances of different assay technology working with customers like Vizgen, TakaraBio, Atrandi. Scientists in 15/20 top market cap pharma, biotech startups and many academic labs use Latch, many of them without realizing its a standalone company. This foundation has proven to be an effective way to think about, engineer and deploy a more powerful tool for scientists: agents for biological data.

English

Because often used as a protein layer on top of more data intensive, standalone measurements, found it more effective to work with spatial vendors on integration rather than direct engagement. ~50-plex antibody panels on their own are not terribly compute intensive. But will eventually change
English

@fruit_struck do not attribute to AI what you can more simply attribute to a tired idiot
English

@kenbwork Do mutations *affect* autism development. I view grammatical errors as proof of human in text but maybe not images. I know historically Latch has published AI hallucinated citations
English

Have learned a lot building and deploying frontier agents into pharma over the past few months. Believe progress in biotech will be faster than many anticipate, and we can learn a lot from how software is unfolding.
It’s unlikely biology will jump straight to fully autonomous AI scientists. Like software, agents first get useful where work is executable, feedback-rich, and economically bottlenecked. In software, that substrate is code. In biology, it is measurement-grounded data analysis.
As agents reliably turn raw molecular data into trusted scientific outputs, they become the interface through which AI starts to understand biology. Essay below:
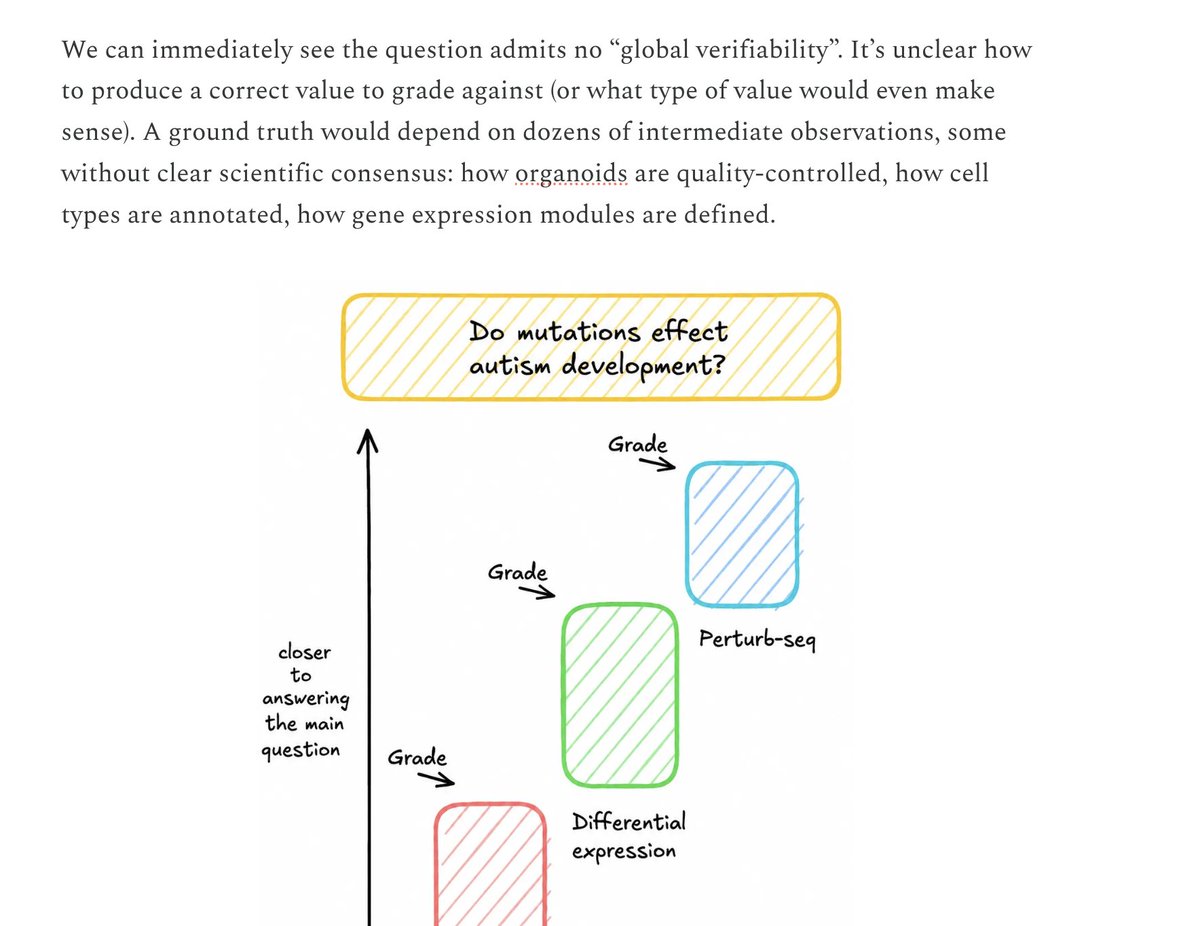
English

Think this essay supports these claims. "Biology has a code-like layer, therefore biology agents will work like software agents" is the obvious analogy and exactly what I'm trying to avoid.
The argument is more: "Software agents prove that executable feedback loops can bootstrap complex capability. Biology can use some of that machinery, especially coding, execution, inspection, debugging, but the scientific loop is structurally different because the objects are experimental measurements, leaky abstractions, and claims about living systems."
English

While I generally agree with the point that agents being good at code means the first application in bio will be agents writing code (to paraphrase), this misses the key point about how to enable this effectively: context.
Kenny Workman@kenbwork
Have learned a lot building and deploying frontier agents into pharma over the past few months. Believe progress in biotech will be faster than many anticipate, and we can learn a lot from how software is unfolding. It’s unlikely biology will jump straight to fully autonomous AI scientists. Like software, agents first get useful where work is executable, feedback-rich, and economically bottlenecked. In software, that substrate is code. In biology, it is measurement-grounded data analysis. As agents reliably turn raw molecular data into trusted scientific outputs, they become the interface through which AI starts to understand biology. Essay below:
English




