Chamindu jayathilake retweetledi

Nearly 200 years after nicotine was first chemically isolated, we’ve finally figured out its complete biosynthesis pathway. Doing so required an insane effort and many years of work. The authors — a Chinese group — ended up crossing 643 lines of tobacco plants to find a single mutant incapable of making nicotine. They next backcrossed and inbred that plant to figure out the specific mutations, in various genes, and map the enzymes responsible.
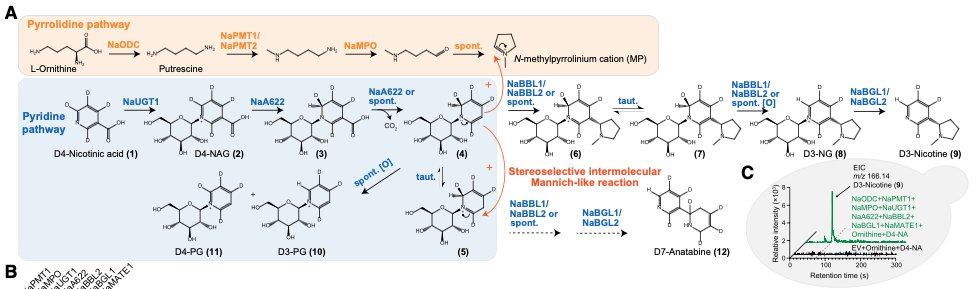
Nicotine is made from two “ring-shaped” molecules fused together. One ring has five carbons (the “pyrrolidine ring”) and the second has six carbons (the “pyridine ring.”) Scientists already knew quite a bit about how these rings get made, but not every step, and not how tthey join together to make nicotine.
The pyrrolidine ring starts when ornithine, an amino acid that is not used to make proteins, gets its carbon dioxide clipped off by an enzyme, called ornithine decarboxylase, to make putrescine. This putrescine then has a methyl group attached to it, and gets oxidized. At this point, the molecule is a chain with four carbon atoms; one end has an amine, and the other a methylated amine. The amine end gets cut off and replaced with a reactive aldehyde; the chain folds into a loop; and the methylated amine “attacks” electrons on the aldehyde to form the ring.
To make the pyridine ring, plant cells first take aspartate (the amino acid) and oxidize it. The resulting molecule is then transformed into nicotinic acid mononucleotide, which is just vitamin B3 with a sugar and phosphate attached. This paper is the first to report that NAMN hydrolase clips off the sugar and phosphate to release pure vitamin B3; also called niacin or nicotinic acid. (The names are slightly confusing.)
The paper’s major contribution, though, is in figuring out how the two rings get fused together. The nicotinic acid is unstable, so an enzyme quickly attaches a sugar to it. Another enzyme, called A622, then strips off a CO2 group, making the molecule reactive again. And finally, that reactive intermediate “attacks” the five-membered pyrrolidine ring to join the two halves together. Other enzymes strip off the remaining sugar to make nicotine. (This whole pathway is shown in the image below.)
All of this happens on the surface of plant vacuoles. Many of the chemical intermediates are toxic, so they need to be sequestered and converted quickly. And as soon as the final nicotine gets made, a transporter pumps it into the vacuole, where it is stored away.
It’s actually difficult to wrap my head around the amount of work packed into this paper, so I’ll just give some quick bullet points:
1. They grew 643 inbred plant lines, which were made by crossing together 26 different parent tobacco plants. They extracted metabolites from all of them.
2. They did a bunch of single-cell RNA sequencing on the tobacco roots to figure out which cells actually express the nicotine biosynthesis genes.
3. “Stumbled” upon a mutant plant which was not able to make nicotine, and then sequenced its entire genome. They also crossed back this plant and inbred it for two generations to find the mutation responsible; a single C-to-T swap. This experiment alone must have taken at least two years of work.
4. Fed plants with isotopically “heavy” nicotinic acid and then tracked its movements through metabolic pathways.
5. Collected at least 630 mass spectrometry spectra.
6. RECONSTITUTED THE ENTIRE PATHWAY IN FOUR DIFFERENT SPECIES: YEAST, TOMATO, EGGPLANTS, AND PEAS (!!!!!!!!)
7. And a lot more…
Anyway, insane paper. China has been putting out incredible plant biology papers for the last several years.

English