Deep Seq
161 posts


Deep Seq
@DeepSeqNotts
Core Next Generation Sequencing facility at the University of Nottingham. Experts in Nanopore, Illumina and Sanger sequencing
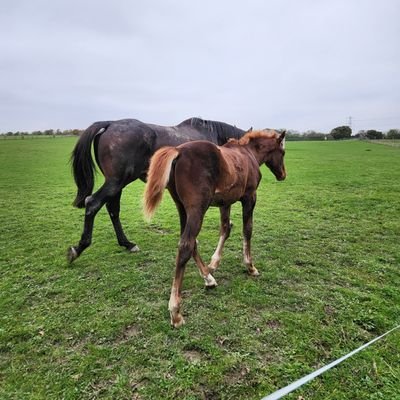


This gif is from one sample of three run on a single @nanopore promethION flow cell. Frankly I'm super impressed at the performance though there is headroom for more improvements. We ran the same libraries as on gridION and have updated our preprint here. biorxiv.org/content/10.110…




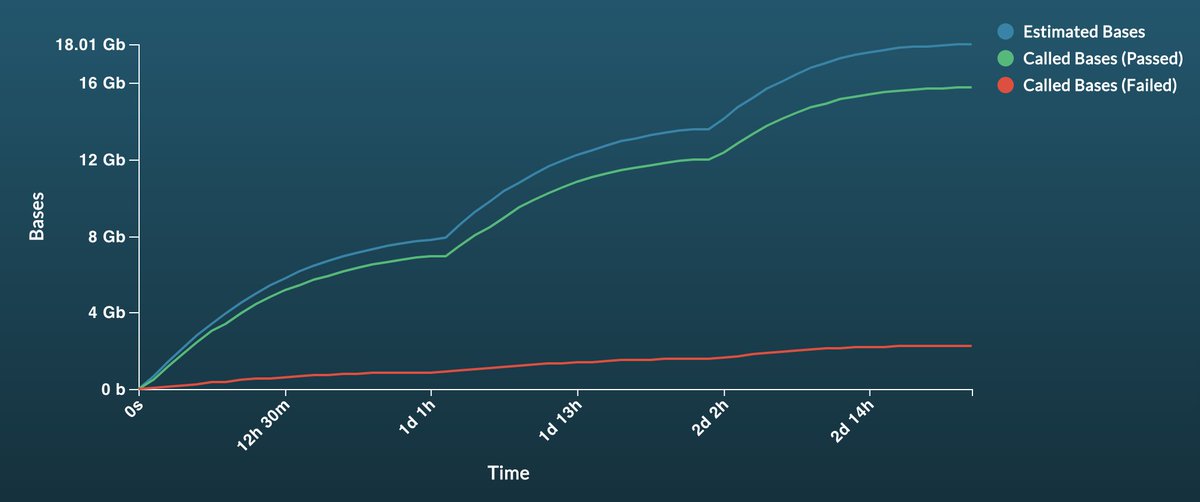
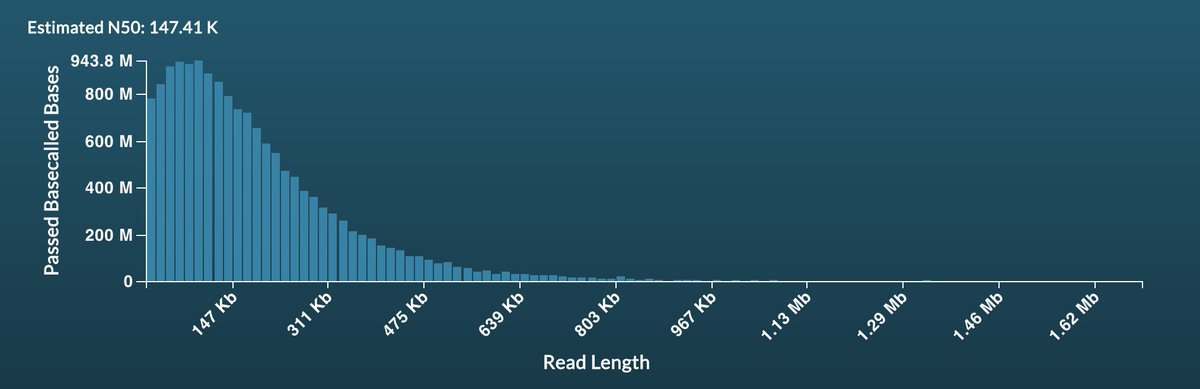



So - our ReadFish paper came out on Monday nature.com/articles/s4158… using GPU - but what if you only have a laptop? Here's an update from @alexomics to our tools that enable ReadFish on Mac, Linux or Windows with no additional GPU or compute. looselab.github.io/2020/12/02/rea… for details.

Great to see this paper out - loads of hard work from @alexomics and many others to get this completed - adaptive sampling on @nanopore sequencers targeting hundreds of genes to depths in excess of 40x on a single flowcell. nature.com/articles/s4158…













