Devin Leopold
35 posts



@pseudotsugonoid fwiw - I never heard about the GG issue you mentioned. I just spot checked some samples from a novaseq 2x250 single 12bp indexed run and there were plenty of reads for samples with GG on either end of the index.
English

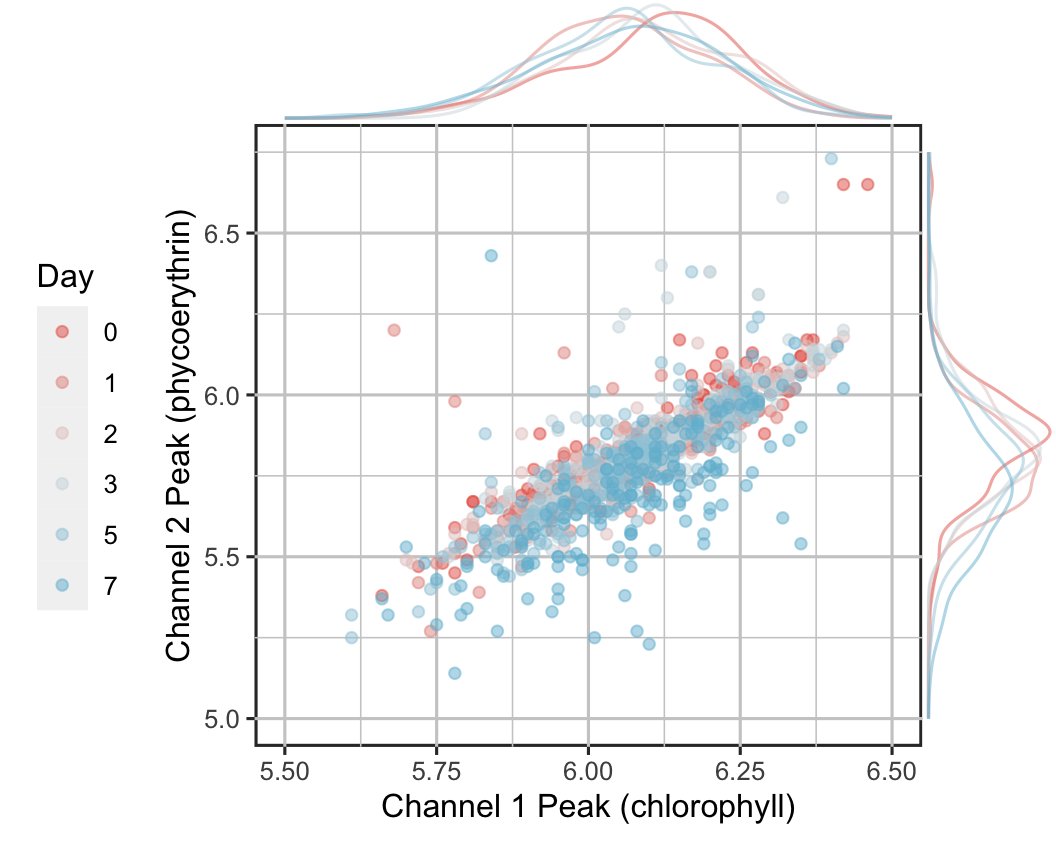
Hi folks--I'd appreciate some wisdom in the area of Illumina 2-channel SBS. My SEQanswers post has it all:
seqanswers.com/forum/sequenci…
Basically, are leading-GG indices unusable? Illumina examples seem to suggest that just having equal base diversity at each position is sufficient...
English

@pseudotsugonoid Is it that the examples in your post are acceptable because the i7 index is always read in reverse?
English

@chesscom_in Qh8 is the only guarantee of mate in 2. Other solutions don't account for black moving Kf1
English

#phish They only scanned one of our tickets on the way into Dicks tonight. Respond if you are outside and need a ticket.
English

@mixotrophe Maybe you could modify the one on the left so the values are shown by the proportion of the box filled in as opposed to the hue/shade. That way you can keep the color scheme but the values would be easier to compare (and interpret).
English

OK sci-twitter, please help again! Which of these 3 do you prefer? Trying to show pathway completeness (rows) for 4 species of #Mesodinium (columns). Box saturation = completeness. (Lead author doesn't think left works b/c hard to compare across colors). #figaday 2.7/n #year2

English

📢 {shinytest2} v0.2.0 is on CRAN! #rstats
🕸️ rstudio.github.io/shinytest2/new…
Breaking:
* Downloaded file names
* Default `load_timeout`: 15s
* Remaining default `timeout`s: 4s
Features:
* Fuzzy screenshot comparison
* Seamless `test_app()` reporter
* Skip test if {chromote} can't start
English

@e_hannula @ldereske @MAnthony02 @PacBio ITSx extracts the synthetic sequences from the amptk paper just fine in my experience. The ITSx model only searches for the conserved regions flanking ITS which are based on real fungal sequences in the amptk synmock seqs.
English

@ldereske @MAnthony02 @PacBio Indeed, you should loose the synthetic sequences if using ITSx (which is a sign it works 😉). I don't use spike in mock but rather selfmade mock community (made of variety of cultures) - and it performs well with ITSx. I can live with relative abundances...
English
Devin Leopold retweetledi

Launch your project into the stratosphere! Retweet this for the chance to win a dream Thelio Major desktop worth up to $10,000! One entry per person, contest ends Sept 30th. Learn more about System76's out-of-this-world desktop computers here! s76.co/LIL-Thelio
#System76

English

@BentonRochester @drewmck The relative abundances we recover from eDNA sequencing are subject to multiple sources of bias, and should be interpreted with that in mind. But your bees were most definitely consuming everything we detected.
English

@drewmck I don’t unfortunately. It’s hard to believe cucumber was a primary flower that they pollinated but we do have a farm a half mile from us and bees go up to three miles to forage.
English

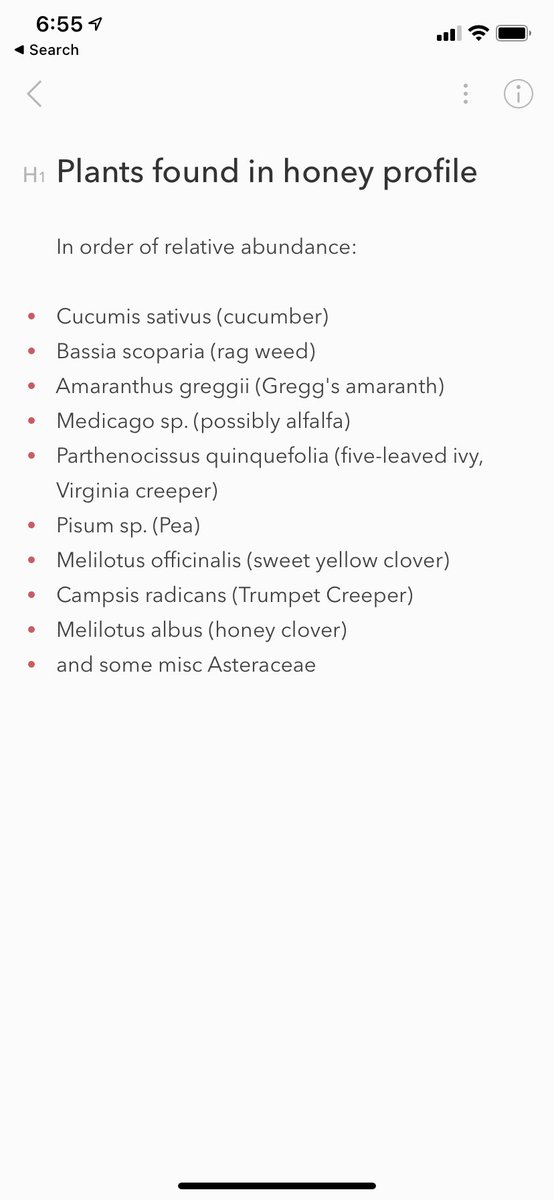
My neighbor profiled the honey my bees produced. These are the plants he found. Science is cool!
#beekeeping

English

@jdierks @BentonRochester This was done by sequencing plant DNA present in the honey, which is one of the eDNA sequencing services we offer @JonahVentures.
English

@BentonRochester How is this done? Mass spectrometry of some sort, maybe?
English
Devin Leopold retweetledi

In this paper, Leopold et al. show that the diversity of putative #ericoid #mycorrhizal #fungi increases with soil age and progressive #phosphorus limitation across a 4.1-million-year #chronosequence
@DevinLeopold
#FEMSMicrobiolEcol #pedogenesis
academic.oup.com/femsec/article…

English
Devin Leopold retweetledi

Devin Leopold is our bioinformatician. All the data you get back from us, whether in files or on the data portal goes through him first. Welcome to @JonahVentures, Devin!

English
Devin Leopold retweetledi

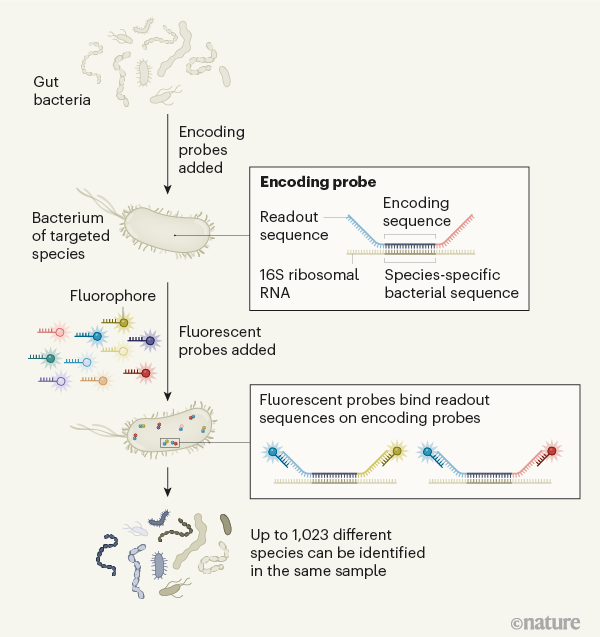
HIPR-FISH: Highly multiplexed spatial mapping of microbial communities. Visualize microbes in the gut with high taxonomic resolution! It was super fun to collaborate with @DeVlaminckLab and @haoiseetheworld on this! @nature @CornellBME

English
Devin Leopold retweetledi

I’m excited to invite applications for two PhD positions in my lab at UT Arlington! Love insects? Love the microbiome? Check it out and spread the word: wfscjobs.tamu.edu/jobs/phd-posit…
English
Devin Leopold retweetledi

hi all! Im looking for a new postdoc on an NSF funded fungal endosymbiont interaction genomics project, see more detail at gradschool.oregonstate.edu/sites/gradscho… please RT
English